-検索条件
-検索結果
検索 (著者・登録者: johnson & ag)の結果156件中、1から50件目までを表示しています

EMDB-41371: 
Cryo-EM structure of the Rev1-Polzeta-DNA-dCTP complex

EMDB-41372: 
Cryo-EM structure of Rev1(deltaN)-Polzeta-DNA-dCTP complex

PDB-8tlq: 
Cryo-EM structure of the Rev1-Polzeta-DNA-dCTP complex

PDB-8tlt: 
Cryo-EM structure of Rev1(deltaN)-Polzeta-DNA-dCTP complex
EMDB-42478: 
Trehalose Synthase (TreS) of Mycobacterium tuberculosis in complex with 6-TreAz compound

PDB-8uqv: 
Trehalose Synthase (TreS) of Mycobacterium tuberculosis in complex with 6-TreAz compound

EMDB-43508: 
Structure of a bacterial gasdermin small oval pore assembly

EMDB-43509: 
Structure of a bacterial gasdermin medium oval pore assembly

EMDB-43510: 
Structure of a bacterial gasdermin large oval pore assembly

EMDB-43511: 
Structure of a bacterial gasdermin double pore assembly
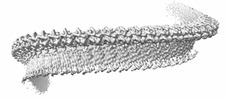

EMDB-43513: 
Structure of a bacterial gasdermin slinky-like oligomer from a heterogeneous sample

EMDB-41370: 
Structure of a class A GPCR/Fab complex

EMDB-41827: 
Structure of a class A GPCR/agonist complex (focused map2)

EMDB-41828: 
Structure of a class A GPCR/agonist complex (focused map1)

EMDB-41829: 
Structure of a class A GPCR/agonist complex

EMDB-41850: 
Structure of a class A GPCR/agonist complex (Consensus map)

PDB-8tlm: 
Structure of a class A GPCR/Fab complex

PDB-8u1u: 
Structure of a class A GPCR/agonist complex

EMDB-41983: 
Structure of the phage immune evasion protein Gad1 bound to the Gabija GajAB complex

EMDB-29068: 
Cryo-EM structure of the GR-Hsp90-FKBP52 complex

EMDB-29069: 
Cryo-EM structure of the GR-Hsp90-FKBP51 complex

PDB-8ffv: 
Cryo-EM structure of the GR-Hsp90-FKBP52 complex

PDB-8ffw: 
Cryo-EM structure of the GR-Hsp90-FKBP51 complex

EMDB-41153: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf

EMDB-41154: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf

PDB-8tcf: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf

PDB-8tcg: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf

EMDB-29930: 
T. cruzi topoisomerase II alpha bound to dsDNA and the covalent inhibitor CT1

PDB-8gcc: 
T. cruzi topoisomerase II alpha bound to dsDNA and the covalent inhibitor CT1

EMDB-15084: 
cryo-EM structure of thioredoxin glutathione reductase in complex with a non-competitive inhibitor

PDB-8a1r: 
cryo-EM structure of thioredoxin glutathione reductase in complex with a non-competitive inhibitor

EMDB-40570: 
Structure of a bacterial gasdermin slinky-like oligomer

EMDB-29323: 
Structure of RdrA from Escherichia coli RADAR defense system

EMDB-29324: 
Map of RdrA from Escherichia coli RADAR defense system in single-split conformation

EMDB-29325: 
Map of RdrA from Escherichia coli RADAR defense system in double-split conformation

EMDB-29326: 
Structure of RdrA from Streptococcus suis RADAR defense system

EMDB-29327: 
Structure of RdrB from Escherichia coli RADAR defense system

EMDB-29328: 
Structure of RdrA-RdrB complex from Escherichia coli RADAR defense system

EMDB-14885: 
OMI-42 FAB IN COMPLEX WITH SARS-COV-2 BETA SPIKE GLYCOPROTEIN

EMDB-14886: 
OMI-38 FAB IN COMPLEX WITH SARS-COV-2 BETA SPIKE RBD (local refinement)

EMDB-14887: 
OMI-2 FAB IN COMPLEX WITH SARS-COV-2 BETA SPIKE GLYCOPROTEIN

EMDB-14910: 
OMI-38 FAB IN COMPLEX WITH SARS-COV-2 BETA SPIKE

PDB-7zr7: 
OMI-42 FAB IN COMPLEX WITH SARS-COV-2 BETA SPIKE GLYCOPROTEIN

PDB-7zr8: 
OMI-38 FAB IN COMPLEX WITH SARS-COV-2 BETA SPIKE RBD (local refinement)

PDB-7zr9: 
OMI-2 FAB IN COMPLEX WITH SARS-COV-2 BETA SPIKE GLYCOPROTEIN

PDB-7zrc: 
OMI-38 FAB IN COMPLEX WITH SARS-COV-2 BETA SPIKE

EMDB-26429: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the neutralizing antibody Fab fragment, C1520

EMDB-26430: 
Structure of the SARS-CoV-2 NTD in complex with C1520, local refinement

EMDB-26431: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the neutralizing antibody Fab fragment, C1717

EMDB-26432: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the neutralizing antibody Fab fragment, C1791
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
