-検索条件
-検索結果
検索 (著者・登録者: chuang & d)の結果293件中、1から50件目までを表示しています

EMDB-36228: 
Cryo-EM structure of Mi3 fused with FKBP

EMDB-36230: 
Cryo-EM structure of Mi3 fused with LOV2

PDB-8jga: 
Cryo-EM structure of Mi3 fused with FKBP

PDB-8jgc: 
Cryo-EM structure of Mi3 fused with LOV2
EMDB-36651: 
hOCT1 in complex with metformin in outward open conformation
EMDB-36652: 
hOCT1 in complex with metformin in outward occluded conformation
EMDB-36653: 
hOCT1 in complex with metformin in inward occluded conformation
EMDB-36654: 
hOCT1 in complex with nb5660 in inward facing partially open 1 conformation
EMDB-36655: 
hOCT1 in complex with nb5660 in inward facing fully open conformation
EMDB-36656: 
hOCT1 in complex with nb5660 in inward facing partially open 2 conformation
EMDB-36657: 
hOCT1 in complex with spironolactone in outward facing partially occluded conformation
EMDB-36658: 
hOCT1 in complex with spironolactone in inward facing occluded conformation

PDB-8jts: 
hOCT1 in complex with metformin in outward open conformation

PDB-8jtt: 
hOCT1 in complex with metformin in outward occluded conformation

PDB-8jtv: 
hOCT1 in complex with metformin in inward occluded conformation

PDB-8jtw: 
hOCT1 in complex with nb5660 in inward facing partially open 1 conformation

PDB-8jtx: 
hOCT1 in complex with nb5660 in inward facing fully open conformation

PDB-8jty: 
hOCT1 in complex with nb5660 in inward facing partially open 2 conformation

PDB-8jtz: 
hOCT1 in complex with spironolactone in outward facing partially occluded conformation

PDB-8ju0: 
hOCT1 in complex with spironolactone in inward facing occluded conformation
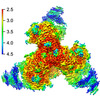
EMDB-28617: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB

EMDB-28618: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-COMBO1 FAB

EMDB-28619: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-MM28 FAB

PDB-8euu: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB

PDB-8euv: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-COMBO1 FAB

PDB-8euw: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-MM28 FAB
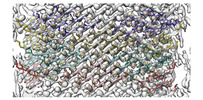

EMDB-27820: 
Near-Atomic Resolution Structure of J-aggregated Helical Light Harvesting Nanotubes

EMDB-34522: 
Cryo-EM Structure of SARS-CoV-2 BA.2 Spike protein in complex with BA7535

EMDB-34526: 
Cryo-EM structure of SARS-CoV-2 BA.2 RBD in complex with BA7535 fab (local refinement)

PDB-8h7l: 
Cryo-EM Structure of SARS-CoV-2 BA.2 Spike protein in complex with BA7535

PDB-8h7z: 
Cryo-EM structure of SARS-CoV-2 BA.2 RBD in complex with BA7535 fab (local refinement)

EMDB-28962: 
LH2-LH3 antenna in antiparallel configuration embedded in a nanodisc

EMDB-28963: 
LH2-LH3 antenna in parallel configuration embedded in a nanodisc

PDB-8fb9: 
LH2-LH3 antenna in anti parallel configuration embedded in a nanodisc

PDB-8fbb: 
LH2-LH3 antenna in parallel configuration embedded in a nanodisc

EMDB-33924: 
Cryo-EM structure of Nb29-alpha1AAR-miniGsq complex bound to oxymetazoline
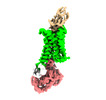
EMDB-33928: 
Cryo-EM structure of Nb29-alpha1AAR-miniGsq complex bound to noradrenaline
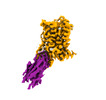
EMDB-33930: 
Cryo-EM structure of alpha1AAR-Nb6 complex bound to tamsulosin

PDB-7ym8: 
Cryo-EM structure of Nb29-alpha1AAR-miniGsq complex bound to oxymetazoline

PDB-7ymh: 
Cryo-EM structure of Nb29-alpha1AAR-miniGsq complex bound to noradrenaline

PDB-7ymj: 
Cryo-EM structure of alpha1AAR-Nb6 complex bound to tamsulosin

EMDB-33632: 
ApoSIDT2-pH7.4

EMDB-33637: 
SIDT2-pH5.5 plus miRNA

EMDB-33638: 
ApoSIDT2-pH5.5

PDB-7y63: 
ApoSIDT2-pH7.4

PDB-7y68: 
SIDT2-pH5.5 plus miRNA

PDB-7y69: 
ApoSIDT2-pH5.5

EMDB-33845: 
Cryo-EM map of Rpd3S complex

EMDB-33846: 
Cryo-EM map of Rpd3S:head-bridge-right arm

EMDB-33847: 
Cryo-EM map of Rpd3S:bridge-left arm
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
