+ Open data
Open data
- Basic information
Basic information
| Entry | 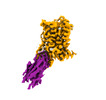 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of alpha1AAR-Nb6 complex bound to tamsulosin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  GPCR / GPCR /  Nanobody / Nanobody /  Antagonist / Antagonist /  Complex / Complex /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Biological species |   Homo sapiens (human) / Homo sapiens (human) /   Lama glama (llama) Lama glama (llama) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.35 Å cryo EM / Resolution: 3.35 Å | |||||||||
 Authors Authors | Toyoda Y / Zhu A / Yan C / Kobilka BK / Liu X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural basis of α-adrenergic receptor activation and recognition by an extracellular nanobody. Authors: Yosuke Toyoda / Angqi Zhu / Fang Kong / Sisi Shan / Jiawei Zhao / Nan Wang / Xiaoou Sun / Linqi Zhang / Chuangye Yan / Brian K Kobilka / Xiangyu Liu /    Abstract: The αadrenergic receptor (αAR) belongs to the family of G protein-coupled receptors that respond to adrenaline and noradrenaline. αAR is involved in smooth muscle contraction and cognitive ...The αadrenergic receptor (αAR) belongs to the family of G protein-coupled receptors that respond to adrenaline and noradrenaline. αAR is involved in smooth muscle contraction and cognitive function. Here, we present three cryo-electron microscopy structures of human αAR bound to the endogenous agonist noradrenaline, its selective agonist oxymetazoline, and the antagonist tamsulosin, with resolutions range from 2.9 Å to 3.5 Å. Our active and inactive αAR structures reveal the activation mechanism and distinct ligand binding modes for noradrenaline compared with other adrenergic receptor subtypes. In addition, we identified a nanobody that preferentially binds to the extracellular vestibule of αAR when bound to the selective agonist oxymetazoline. These results should facilitate the design of more selective therapeutic drugs targeting both orthosteric and allosteric sites in this receptor family. | |||||||||
| History |
|
- Structure visualization
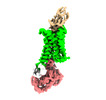
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33930.map.gz emd_33930.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33930-v30.xml emd-33930-v30.xml emd-33930.xml emd-33930.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33930.png emd_33930.png | 55.7 KB | ||
| Others |  emd_33930_half_map_1.map.gz emd_33930_half_map_1.map.gz emd_33930_half_map_2.map.gz emd_33930_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33930 http://ftp.pdbj.org/pub/emdb/structures/EMD-33930 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33930 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33930 | HTTPS FTP |
-Related structure data
| Related structure data |  7ymjMC  7ym8C  7ymhC C: citing same article ( M: atomic model generated by this map |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33930.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33930.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_33930_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_33930_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : alpha1AAR-Nb6 complex
| Entire | Name: alpha1AAR-Nb6 complex |
|---|---|
| Components |
|
-Supramolecule #1: alpha1AAR-Nb6 complex
| Supramolecule | Name: alpha1AAR-Nb6 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|
-Supramolecule #2: alpha1AAR
| Supramolecule | Name: alpha1AAR / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Nb6
| Supramolecule | Name: Nb6 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:   Lama glama (llama) Lama glama (llama) |
-Macromolecule #1: alpha1A-adrenergic receptor
| Macromolecule | Name: alpha1A-adrenergic receptor / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.80418 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDAMVFLSG QASDSSQCTQ PPAPVQISKA ILLGVILGGL ILFGVLGNIL VILSVACHRH LHSVTHYYI VNLAVADLLL TSTVLPFSAI FEVLGYWAFG RVFCNIWAAV DVLCCTARIW GLCIISIDRY IGVSYPLRYP T IVTQRRGL ...String: MKTIIALSYI FCLVFADYKD DDDAMVFLSG QASDSSQCTQ PPAPVQISKA ILLGVILGGL ILFGVLGNIL VILSVACHRH LHSVTHYYI VNLAVADLLL TSTVLPFSAI FEVLGYWAFG RVFCNIWAAV DVLCCTARIW GLCIISIDRY IGVSYPLRYP T IVTQRRGL MALLCVWALS LVISIGPLFG WRQPAPEDET ICQINEEPGY VLFSALGSFY LPLAIILVMY TLMILRLKSV RL LSGSREK DRNLRRITRL VLIVVGCFVL CWLPFFLVMP IGSFFPDFKP SETVFKIVFW LGYLNSCINP IIYPCSSQEF KKA FQNVLR IQCLCRKQSS KHALGYTLHP PSQAVEGQHH HHHHHH |
-Macromolecule #2: Nb6
| Macromolecule | Name: Nb6 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Lama glama (llama) Lama glama (llama) |
| Molecular weight | Theoretical: 16.51559 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MKYLLPTAAA GLLLLAAQPA MAMAQVQLQE SGGGLVQAGE SLRLSCAASG TIFRLYDMGW YRRVSGNQRE LVASITSGGS TKYGDSVKG RFTISRDNAK NTVYLQMSSL KPEDTAVYYC NAEYRTGIWE ELLDGWGQGT QVTVSSHHHH HH |
-Macromolecule #3: Tamsulosin
| Macromolecule | Name: Tamsulosin / type: ligand / ID: 3 / Number of copies: 1 / Formula: JGX |
|---|---|
| Molecular weight | Theoretical: 408.512 Da |
| Chemical component information |  ChemComp-JGX: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.3 µm Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.3 µm |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.35 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 285000 |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X

















