-Search query
-Search result
Showing 1 - 50 of 137 items for (author: grunewald & k)

EMDB-19837: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

EMDB-19838: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state active site with 1-bp DNA mismatch
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

EMDB-19839: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch consensus map
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

EMDB-19840: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch catalytic core focused map
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

EMDB-19841: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch processivity factor focused refinement
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

PDB-9enp: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

PDB-9enq: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state active site with 1-bp DNA mismatch
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM
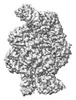
EMDB-18963: 
Structure of the SFTSV L protein in a transcription-priming state without capped RNA [TRANSCRIPTION-PRIMING (in vitro)]
Method: single particle / : Williams HM, Thorkelsson SR, Vogel D, Busch C, Milewski M, Cusack S, Grunewald K, Quemin ERJ, Rosenthal M

EMDB-18967: 
Structure of the SFTSV L protein in a transcription-priming state with bound capped RNA [TRANSCRIPTION-PRIMING]
Method: single particle / : Williams HM, Thorkelsson SR, Vogel D, Busch C, Milewski M, Cusack S, Grunewald K, Quemin ERJ, Rosenthal M

EMDB-18969: 
Structure of the SFTSV L protein stalled in a transcription-specific early elongation state with bound capped RNA [TRANSCRIPTION-EARLY-ELONGATION]
Method: single particle / : Williams HM, Thorkelsson SR, Vogel D, Busch C, Milewski M, Cusack S, Grunewald K, Quemin ERJ, Rosenthal M

PDB-8r6u: 
Structure of the SFTSV L protein in a transcription-priming state without capped RNA [TRANSCRIPTION-PRIMING (in vitro)]
Method: single particle / : Williams HM, Thorkelsson SR, Vogel D, Busch C, Milewski M, Cusack S, Grunewald K, Quemin ERJ, Rosenthal M

PDB-8r6w: 
Structure of the SFTSV L protein in a transcription-priming state with bound capped RNA [TRANSCRIPTION-PRIMING]
Method: single particle / : Williams HM, Thorkelsson SR, Vogel D, Busch C, Milewski M, Cusack S, Grunewald K, Quemin ERJ, Rosenthal M

PDB-8r6y: 
Structure of the SFTSV L protein stalled in a transcription-specific early elongation state with bound capped RNA [TRANSCRIPTION-EARLY-ELONGATION]
Method: single particle / : Williams HM, Thorkelsson SR, Vogel D, Busch C, Milewski M, Cusack S, Grunewald K, Quemin ERJ, Rosenthal M

EMDB-18482: 
Herpes simplex virus 1 capsid (WT) vertices in perinuclear NEC-coated vesicles determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D

EMDB-18484: 
Herpes simplex virus 1 nuclear egress complex (WT) determined in situ from perinuclear vesicles
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D

EMDB-17013: 
HSV-1 DNA polymerase-processivity factor complex in halted elongation state consensus map
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-17014: 
Consensus map of HSV-1 DNA polymerase-processivity factor complex in pre-translocation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-17018: 
Consensus map of HSV-1 DNA polymerase-processivity factor complex in exonuclease state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-17974: 
Pseudorabies virus cytosolic C-capsid (US3 KO) vertices determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-17975: 
Pseudorabies virus primary enveloped (perinuclear) C-capsid (US3 KO) vertices determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-17976: 
Pseudorabies nuclear C-capsids (US3 KO) vertices determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-18473: 
Subtomogram average of pseudorabies virus nuclear egress complex helical form (UL31/34) determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-18474: 
Subtomogram average of pseudorabies virus nuclear egress complex (UL31/34) determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-18479: 
Pseudorabies virus cytosolic C-capsid (WT) vertices determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D

EMDB-18480: 
Pseudorabies virus nuclear C-capsid (WT) vertices determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D
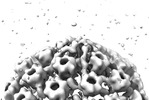

EMDB-18481: 
Herpes simplex virus 1 cytosolic C-capsid (WT) vertices determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D
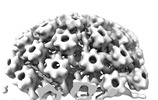
EMDB-18483: 
Herpes simplex virus 1 nuclear C-capsid (WT) vertices determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D

EMDB-16918: 
Focused refinement map of HSV-1 DNA polymerase in pre-translocation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-16919: 
Focused refinement map of HSV-1 DNA polymerase processivity factor in pre-translocation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-16924: 
Focused refinement of HSV-1 DNA polymerase in halted elongation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-16925: 
Focused refinement of HSV-1 DNA polymerase processivity factor in halted elongation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-16927: 
Focused refinement map of HSV-1 DNA polymerase in exonuclease state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-16928: 
Focused refinement map of HSV-1 DNA polymerase processivity factor in exonuclease state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-16906: 
HSV-1 DNA polymerase-processivity factor complex in pre-translocation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg M

EMDB-16907: 
HSV-1 DNA polymerase-processivity factor complex in halted elongation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

EMDB-16909: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM
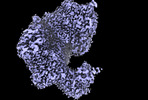
EMDB-16910: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state active site
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

EMDB-16911: 
HSV-1 DNA polymerase active site in alternative exonuclease state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

EMDB-16912: 
HSV-1 DNA polymerase beta-hairpin loop
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

PDB-8oj6: 
HSV-1 DNA polymerase-processivity factor complex in pre-translocation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

PDB-8oj7: 
HSV-1 DNA polymerase-processivity factor complex in halted elongation state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

PDB-8oja: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

PDB-8ojb: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state active site
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

PDB-8ojc: 
HSV-1 DNA polymerase active site in alternative exonuclease state
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

PDB-8ojd: 
HSV-1 DNA polymerase beta-hairpin loop
Method: single particle / : Gustavsson E, Grunewald K, Elias P, Hallberg BM

EMDB-17704: 
Subtomogram average of Vaccinia A10 trimer with open center from in vitro cores
Method: subtomogram averaging / : Turonova B, Liu J

EMDB-17708: 
Subtomogram average of Vaccinia A10 trimer with tight center from in vitro cores
Method: subtomogram averaging / : Turonova B, Liu J

EMDB-17753: 
Subtomogram average of Vaccinia A10 trimer from in situ cores
Method: subtomogram averaging / : Turonova B, Liu J

EMDB-42250: 
Varicella-zoster virus glycoprotein B; H527P prefusion class I.
Method: subtomogram averaging / : Zhou M, Oliver SL, Muyuan C, Vollmer B

EMDB-42251: 
Varicella-zoster virus glycoprotein B; H527P prefusion mutant class II.
Method: subtomogram averaging / : Zhou M, Oliver SL, Muyuan C, Vollmer B
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
