[English] 日本語
 Yorodumi
Yorodumi- EMDB-44537: Human DNA polymerase theta helicase domain dimer bound to DNA in ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human DNA polymerase theta helicase domain dimer bound to DNA in the microhomology aligning conformation | |||||||||
 Map data Map data | Map locally filtered by resolution | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  DNA repair / TMEJ / MMEJ / DNA repair / TMEJ / MMEJ /  DNA binding protein DNA binding protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationsingle-stranded DNA endodeoxyribonuclease activity / double-strand break repair via alternative nonhomologous end joining / HDR through MMEJ (alt-NHEJ) / single-stranded DNA helicase activity / negative regulation of double-strand break repair via homologous recombination / replication fork processing / site of DNA damage / 5'-deoxyribose-5-phosphate lyase activity / somatic hypermutation of immunoglobulin genes / error-prone translesion synthesis ...single-stranded DNA endodeoxyribonuclease activity / double-strand break repair via alternative nonhomologous end joining / HDR through MMEJ (alt-NHEJ) / single-stranded DNA helicase activity / negative regulation of double-strand break repair via homologous recombination / replication fork processing / site of DNA damage / 5'-deoxyribose-5-phosphate lyase activity / somatic hypermutation of immunoglobulin genes / error-prone translesion synthesis /  DNA helicase activity / DNA helicase activity /  base-excision repair / protein homooligomerization / base-excision repair / protein homooligomerization /  RNA-directed DNA polymerase / RNA-directed DNA polymerase /  RNA-directed DNA polymerase activity / double-strand break repair / site of double-strand break / RNA-directed DNA polymerase activity / double-strand break repair / site of double-strand break /  DNA helicase / damaged DNA binding / DNA helicase / damaged DNA binding /  DNA-directed DNA polymerase / DNA-directed DNA polymerase /  DNA-directed DNA polymerase activity / DNA-directed DNA polymerase activity /  DNA repair / DNA damage response / DNA repair / DNA damage response /  chromatin binding / chromatin binding /  Golgi apparatus / magnesium ion binding / Golgi apparatus / magnesium ion binding /  ATP hydrolysis activity / ATP hydrolysis activity /  nucleoplasm / nucleoplasm /  ATP binding / identical protein binding / ATP binding / identical protein binding /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.5 Å cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Zerio CJ / Lander GC | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Human polymerase theta helicase positions DNA microhomologies for double-strand break repair Authors: Zerio CJ / Bai Y / Sosa-Alvarado BA / Guzi T / Lander GC | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44537.map.gz emd_44537.map.gz | 5.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44537-v30.xml emd-44537-v30.xml emd-44537.xml emd-44537.xml | 25.4 KB 25.4 KB | Display Display |  EMDB header EMDB header |
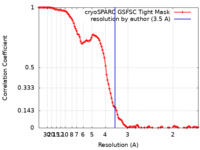
| FSC (resolution estimation) |  emd_44537_fsc.xml emd_44537_fsc.xml | 12.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_44537.png emd_44537.png | 95.2 KB | ||
| Masks |  emd_44537_msk_1.map emd_44537_msk_1.map | 129.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-44537.cif.gz emd-44537.cif.gz | 7.7 KB | ||
| Others |  emd_44537_additional_1.map.gz emd_44537_additional_1.map.gz emd_44537_half_map_1.map.gz emd_44537_half_map_1.map.gz emd_44537_half_map_2.map.gz emd_44537_half_map_2.map.gz | 122.5 MB 120.5 MB 120.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44537 http://ftp.pdbj.org/pub/emdb/structures/EMD-44537 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44537 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44537 | HTTPS FTP |
-Related structure data
| Related structure data |  9bh9MC  9bh6C  9bh7C  9bh8C  9bhaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_44537.map.gz / Format: CCP4 / Size: 129.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44537.map.gz / Format: CCP4 / Size: 129.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map locally filtered by resolution | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.833 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_44537_msk_1.map emd_44537_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Sharp map
| File | emd_44537_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharp map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_44537_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_44537_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human DNA polymerase theta helicase domain bound to stem-loop DNA...
| Entire | Name: Human DNA polymerase theta helicase domain bound to stem-loop DNA with microhomology in the 3' overhang |
|---|---|
| Components |
|
-Supramolecule #1: Human DNA polymerase theta helicase domain bound to stem-loop DNA...
| Supramolecule | Name: Human DNA polymerase theta helicase domain bound to stem-loop DNA with microhomology in the 3' overhang type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 20 KDa |
-Supramolecule #2: Human DNA polymerase theta helicase domain
| Supramolecule | Name: Human DNA polymerase theta helicase domain / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Stem-loop DNA with microhomology in the 3' overhang
| Supramolecule | Name: Stem-loop DNA with microhomology in the 3' overhang / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: DNA polymerase theta
| Macromolecule | Name: DNA polymerase theta / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number:  DNA helicase DNA helicase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 99.671344 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: NLLRRSGKRR RSESGSDSFS GSGGDSSASP QFLSGSVLSP PPGLGRCLKA AAAGECKPTV PDYERDKLLL ANWGLPKAVL EKYHSFGVK KMFEWQAECL LLGQVLEGKN LVYSAPTSAG KTLVAELLIL KRVLEMRKKA LFILPFVSVA KEKKYYLQSL F QEVGIKVD ...String: NLLRRSGKRR RSESGSDSFS GSGGDSSASP QFLSGSVLSP PPGLGRCLKA AAAGECKPTV PDYERDKLLL ANWGLPKAVL EKYHSFGVK KMFEWQAECL LLGQVLEGKN LVYSAPTSAG KTLVAELLIL KRVLEMRKKA LFILPFVSVA KEKKYYLQSL F QEVGIKVD GYMGSTSPSR HFSSLDIAVC TIERANGLIN RLIEENKMDL LGMVVVDELH MLGDSHRGYL LELLLTKICY IT RKSASCQ ADLASSLSNA VQIVGMSATL PNLELVASWL NAELYHTDFR PVPLLESVKV GNSIYDSSMK LVREFEPMLQ VKG DEDHVV SLCYETICDN HSVLLFCPSK KWCEKLADII AREFYNLHHQ AEGLVKPSEC PPVILEQKEL LEVMDQLRRL PSGL DSVLQ KTVPWGVAFH HAGLTFEERD IIEGAFRQGL IRVLAATSTL SSGVNLPARR VIIRTPIFGG RPLDILTYKQ MVGRA GRKG VDTVGESILI CKNSEKSKGI ALLQGSLKPV RSCLQRREGE EVTGSMIRAI LEIIVGGVAS TSQDMHTYAA CTFLAA SMK EGKQGIQRNQ ESVQLGAIEA CVMWLLENEF IQSTEASDGT EGKVYHPTHL GSATLSSSLS PADTLDIFAD LQRAMKG FV LENDLHILYL VTPMFEDWTT IDWYRFFCLW EKLPTSMKRV AELVGVEEGF LARCVKGKVV ARTERQHRQM AIHKRFFT S LVLLDLISEV PLREINQKYG CNRGQIQSLQ QSAAVYAGMI TVFSNRLGWH NMELLLSQFQ KRLTFGIQRE LCDLVRVSL LNAQRARVLY ASGFHTVADL ARANIVEVEV ILKNAVPFKS ARKAVDEEEE AVEERRNMRT IWVTGRKGLT EREAAALIVE EARMILQQD LVEM UniProtKB:  DNA polymerase theta DNA polymerase theta |
-Macromolecule #2: Stem-loop DNA with microhomology in the 3' overhang
| Macromolecule | Name: Stem-loop DNA with microhomology in the 3' overhang / type: dna / ID: 2 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 17.264012 KDa |
| Sequence | String: (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DC)(DA)(DC)(DT) (DG)(DT)(DG)(DA)(DG)(DC)(DT)(DT)(DA) (DG)(DC)(DG)(DT)(DT)(DA)(DG)(DA)(DG)(DT) (DA) (DG)(DG)(DT)(DT)(DT)(DT) ...String: (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DC)(DA)(DC)(DT) (DG)(DT)(DG)(DA)(DG)(DC)(DT)(DT)(DA) (DG)(DC)(DG)(DT)(DT)(DA)(DG)(DA)(DG)(DT) (DA) (DG)(DG)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DG)(DC)(DC)(DC)(DG)(DG)(DG) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.25 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. Details: Grids were glow discharged under vacuum for 30 s at 15 mA in a Pelco easiGlow 91000 Glow Discharge Cleaning System. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: HOMEMADE PLUNGER Details: 3 microliters of sample was applied to the surface of the grid, blotted for 2 seconds after the liquid spot on the filter paper stopped spreading, and the grid was plunged into a liquid ...Details: 3 microliters of sample was applied to the surface of the grid, blotted for 2 seconds after the liquid spot on the filter paper stopped spreading, and the grid was plunged into a liquid ethane bath cooled by liquid nitrogen.. | ||||||||||||
| Details | We added 2X pre-annealed DNA to the protein, lowered the NaCl concentration to 100 mM, and purified the DNA-bound complex by SEC. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 150.0 µm / Calibrated defocus max: 2.5 µm / Calibrated defocus min: 0.3 µm / Calibrated magnification: 60024 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Specialist optics | Energy filter - Slit width: 20 eV |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Temperature | Min: 70.0 K / Max: 77.0 K |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 12440 / Average exposure time: 1.4 sec. / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Details | We modeled DNA bases into sharpened density when possible. We modeled the DNA backbone into locally filtered density in low resolution areas. | |||||||||
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 67.19 / Target criteria: Cross-correlation coefficient | |||||||||
| Output model |  PDB-9bh9: |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X