+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of rabbit Slo1 in complex with gamma1/LRRC26 | |||||||||
 Map data Map data | Map sharpened by Phenix | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Ion channel / potassium transport / Ion channel / potassium transport /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmonoatomic ion-gated channel activity / monoatomic ion channel complex /  potassium channel activity / endoplasmic reticulum membrane / potassium channel activity / endoplasmic reticulum membrane /  metal ion binding / metal ion binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) | |||||||||
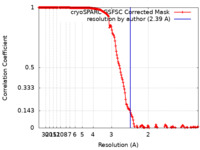
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.39 Å cryo EM / Resolution: 2.39 Å | |||||||||
 Authors Authors | Redhardt M / Raunser S / Raisch T | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: FEBS Lett / Year: 2024 Journal: FEBS Lett / Year: 2024Title: Cryo-EM structure of the Slo1 potassium channel with the auxiliary γ1 subunit suggests a mechanism for depolarization-independent activation. Authors: Milena Redhardt / Stefan Raunser / Tobias Raisch /  Abstract: Mammalian Ca-dependent Slo K channels can stably associate with auxiliary γ subunits which fundamentally alter their behavior. By a so far unknown mechanism, the four γ subunits reduce the need for ...Mammalian Ca-dependent Slo K channels can stably associate with auxiliary γ subunits which fundamentally alter their behavior. By a so far unknown mechanism, the four γ subunits reduce the need for voltage-dependent activation and, thereby, allow Slo to open independently of an action potential. Here, using cryo-EM, we reveal how the transmembrane helix of γ1/LRRC26 binds and presumably stabilizes the activated voltage-sensor domain of Slo1. The activation is further enhanced by an intracellular polybasic stretch which locally changes the charge gradient across the membrane. Our data provide a possible explanation for Slo1 regulation by the four γ subunits and also their different activation efficiencies. This suggests a novel activation mechanism of voltage-gated ion channels by auxiliary subunits. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19691.map.gz emd_19691.map.gz | 400.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19691-v30.xml emd-19691-v30.xml emd-19691.xml emd-19691.xml | 21.4 KB 21.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19691_fsc.xml emd_19691_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_19691.png emd_19691.png | 188 KB | ||
| Filedesc metadata |  emd-19691.cif.gz emd-19691.cif.gz | 6.6 KB | ||
| Others |  emd_19691_additional_1.map.gz emd_19691_additional_1.map.gz emd_19691_additional_2.map.gz emd_19691_additional_2.map.gz emd_19691_half_map_1.map.gz emd_19691_half_map_1.map.gz emd_19691_half_map_2.map.gz emd_19691_half_map_2.map.gz | 249.7 MB 432.4 MB 475.2 MB 475.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19691 http://ftp.pdbj.org/pub/emdb/structures/EMD-19691 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19691 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19691 | HTTPS FTP |
-Related structure data
| Related structure data |  8s3eMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19691.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19691.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map sharpened by Phenix | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.68 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unfiltered map from cryoSPARC
| File | emd_19691_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered map from cryoSPARC | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Map sharpened using DeepEMhancer
| File | emd_19691_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map sharpened using DeepEMhancer | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_19691_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_19691_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Slo1-gamma1 complex
| Entire | Name: Slo1-gamma1 complex |
|---|---|
| Components |
|
-Supramolecule #1: Slo1-gamma1 complex
| Supramolecule | Name: Slo1-gamma1 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) |
-Macromolecule #1: Calcium-activated potassium channel subunit alpha-1
| Macromolecule | Name: Calcium-activated potassium channel subunit alpha-1 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) |
| Molecular weight | Theoretical: 126.029523 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDALIIPVTM EVPCDSRGQR MWWAFLASSM VTFFGGLFII LLWRTLKYLW TVCCHCGGKA KEAQKINNGS SQADGTLKPV DEKEEAVAA EVGWMTSVKD WAGVMISAQT LTGRVLVVLV FALSIGALVI YFIDSSNPIE SCQNFYKDFT LQIDMAFNVF F LLYFGLRF ...String: MDALIIPVTM EVPCDSRGQR MWWAFLASSM VTFFGGLFII LLWRTLKYLW TVCCHCGGKA KEAQKINNGS SQADGTLKPV DEKEEAVAA EVGWMTSVKD WAGVMISAQT LTGRVLVVLV FALSIGALVI YFIDSSNPIE SCQNFYKDFT LQIDMAFNVF F LLYFGLRF IAANDKLWFW LEVNSVVDFF TVPPVFVSVY LNRSWLGLRF LRALRLIQFS EILQFLNILK TSNSIKLVNL LS IFISTWL TAAGFIHLVE NSGDPWENFQ NNQALTYWEC VYLLMVTMST VGYGDVYAKT TLGRLFMVFF ILGGLAMFAS YVP EIIELI GNRKKYGGSY SAVSGRKHIV VCGHITLESV SNFLKDFLHK DRDDVNVEIV FLHNISPNLE LEALFKRHFT QVEF YQGSV LNPHDLARVK IESADACLIL ANKYCADPDA EDASNIMRVI SIKNYHPKIR IITQMLQYHN KAHLLNIPSW NWKEG DDAI CLAELKLGFI AQSCLAQGLS TMLANLFSMR SFIKIEEDTW QKYYLEGVSN EMYTEYLSSA FVGLSFPTVC ELCFVK LKL LMIAIEYKSA NRESRILINP GNHLKIQEGT LGFFIASDAK EVKRAFFYCK ACHDDITDPK RIKKCGCKRL EDEQPST LS PKKKQRNGGM RNSPNSSPKL MRHDPLLIPG NDQIDNMDSN VKKYDSTGMF HWCAPKEIEK VILTRSEAAM TVLSGHVV V CIFGDVSSAL IGLRNLVMPL RASNFHYHEL KHIVFVGSIE YLKREWETLH NFPKVSILPG TPLSRADLRA VNINLCDMC VILSANQNNI DDTSLQDKEC ILASLNIKSM QFDDSIGVLQ ANSQGFTPPG MDRSSPDNSP VHGMLRQPSI TTGVNIPIIT ELVNDTNVQ FLDQDDDDDP DTELYLTQPF ACGTAFAVSV LDSLMSATYF NDNILTLIRT LVTGGATPEL EALIAEENAL R GGYSTPQT LANRDRCRVA QLALLDGPFA DLGDGGCYGD LFCKALKTYN MLCFGIYRLR DAHLSTPSQC TKRYVITNPP YE FELVPTD LIFCLMQFDH NAGQSRASLS HSSHSSQSSS KKSSSVHSIP STANRQNRPK SRESRDKQKY VQEERLLEVL FQ UniProtKB:  Calcium-activated potassium channel subunit alpha-1 Calcium-activated potassium channel subunit alpha-1 |
-Macromolecule #2: Leucine-rich repeat-containing protein 26
| Macromolecule | Name: Leucine-rich repeat-containing protein 26 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) |
| Molecular weight | Theoretical: 35.938766 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MRSPSFSSWP PPLLLLLLLH PWQVCAQAPS LATSSGVSGA PDCPEACACA PGGQANCSAL ALSAVPAGLS RRVRALLLDH NRLGSLPPG AFADADALLR LDLRENELRW VHARAFWGLG ALQRLDLSAN RLEALAPGTF GPLRALRTLS LAGNRLARLE P AALGALPL ...String: MRSPSFSSWP PPLLLLLLLH PWQVCAQAPS LATSSGVSGA PDCPEACACA PGGQANCSAL ALSAVPAGLS RRVRALLLDH NRLGSLPPG AFADADALLR LDLRENELRW VHARAFWGLG ALQRLDLSAN RLEALAPGTF GPLRALRTLS LAGNRLARLE P AALGALPL LRALSLQDNA LSALSPGLLA GLPALDALRL RGNPWTCDCA LRPLCTWLRR HPRPASEAET PVCVSPGRLA RS PLAAFPD AAFRHCARPL SPRDLAMIYL LGPASFLASL VACLALGSAL TACRARRRRR QRTAAHRPPR RSLDLDPGGP ASP ANAGSP AEAGLGRPLE VLFQ |
-Macromolecule #3: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 3 / Number of copies: 16 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #4: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 4 / Number of copies: 8 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #5: (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-...
| Macromolecule | Name: (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSAN-1-AMINIUM 4-OXIDE type: ligand / ID: 5 / Number of copies: 36 / Formula: 6PL |
|---|---|
| Molecular weight | Theoretical: 763.1 Da |
| Chemical component information |  ChemComp-6PL: |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #7: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 7 / Number of copies: 4 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Macromolecule #8: water
| Macromolecule | Name: water / type: ligand / ID: 8 / Number of copies: 13 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.2 µm Bright-field microscopy / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.2 µm |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 52.3 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X