[English] 日本語
 Yorodumi
Yorodumi- EMDB-17836: Consensus cryo-EM structure of Dynein-dynactin-JIP3(1-560)-LIS1 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Consensus cryo-EM structure of Dynein-dynactin-JIP3(1-560)-LIS1 | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords |  Dynein / Dynein /  AAA-Atpase / p150 / AAA-Atpase / p150 /  LIS1 / LIS1 /  MOTOR PROTEIN / MOTOR PROTEIN /  Dynactin / JIP3 Dynactin / JIP3 | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
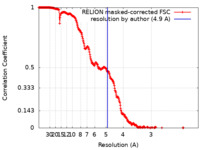
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 4.9 Å cryo EM / Resolution: 4.9 Å | ||||||||||||
 Authors Authors | Singh K / Lau CK / Manigrasso G / Gassmann R / Carter AP | ||||||||||||
| Funding support |  United Kingdom, European Union, 3 items United Kingdom, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: Science / Year: 2024 Journal: Science / Year: 2024Title: Molecular mechanism of dynein-dynactin complex assembly by LIS1. Authors: Kashish Singh / Clinton K Lau / Giulia Manigrasso / José B Gama / Reto Gassmann / Andrew P Carter /   Abstract: Cytoplasmic dynein is a microtubule motor vital for cellular organization and division. It functions as a ~4-megadalton complex containing its cofactor dynactin and a cargo-specific coiled-coil ...Cytoplasmic dynein is a microtubule motor vital for cellular organization and division. It functions as a ~4-megadalton complex containing its cofactor dynactin and a cargo-specific coiled-coil adaptor. However, how dynein and dynactin recognize diverse adaptors, how they interact with each other during complex formation, and the role of critical regulators such as lissencephaly-1 (LIS1) protein (LIS1) remain unclear. In this study, we determined the cryo-electron microscopy structure of dynein-dynactin on microtubules with LIS1 and the lysosomal adaptor JIP3. This structure reveals the molecular basis of interactions occurring during dynein activation. We show how JIP3 activates dynein despite its atypical architecture. Unexpectedly, LIS1 binds dynactin's p150 subunit, tethering it along the length of dynein. Our data suggest that LIS1 and p150 constrain dynein-dynactin to ensure efficient complex formation. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17836.map.gz emd_17836.map.gz | 1.6 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17836-v30.xml emd-17836-v30.xml emd-17836.xml emd-17836.xml | 17.9 KB 17.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17836_fsc.xml emd_17836_fsc.xml | 28.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_17836.png emd_17836.png | 60.4 KB | ||
| Masks |  emd_17836_msk_1.map emd_17836_msk_1.map | 1.9 GB |  Mask map Mask map | |
| Filedesc metadata |  emd-17836.cif.gz emd-17836.cif.gz | 4.2 KB | ||
| Others |  emd_17836_additional_1.map.gz emd_17836_additional_1.map.gz emd_17836_half_map_1.map.gz emd_17836_half_map_1.map.gz emd_17836_half_map_2.map.gz emd_17836_half_map_2.map.gz | 1.8 GB 1.6 GB 1.6 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17836 http://ftp.pdbj.org/pub/emdb/structures/EMD-17836 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17836 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17836 | HTTPS FTP |
-Related structure data
| Related structure data |  8pqvC  8pqwC  8pqyC  8pqzC  8pr0C  8pr1C  8pr2C  8pr3C  8pr4C  8pr5C  8ptkC C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17836.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17836.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.059 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17836_msk_1.map emd_17836_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
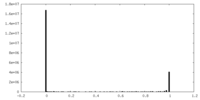
| Density Histograms |
-Additional map: Postprocessed map
| File | emd_17836_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17836_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17836_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Composite structure of Dynein-Dynactin-JIP3-LIS1
| Entire | Name: Composite structure of Dynein-Dynactin-JIP3-LIS1 |
|---|---|
| Components |
|
-Supramolecule #1: Composite structure of Dynein-Dynactin-JIP3-LIS1
| Supramolecule | Name: Composite structure of Dynein-Dynactin-JIP3-LIS1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#7, #9, #11, #10, #12, #8, #13-#19 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.5 µm Bright-field microscopy / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.5 µm |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 53.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Software | Name:  UCSF Chimera UCSF Chimera |
 Movie
Movie Controller
Controller














 Z
Z Y
Y X
X