+Search query
-Structure paper
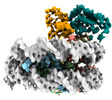
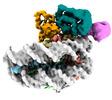
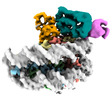
| Title | Mechanisms of RNF168 nucleosome recognition and ubiquitylation. |
|---|---|
| Journal, issue, pages | Mol Cell, Vol. 84, Issue 5, Page 839-853.e12, Year 2024 |
| Publish date | Mar 7, 2024 |
 Authors Authors | Qi Hu / Debiao Zhao / Gaofeng Cui / Janarjan Bhandari / James R Thompson / Maria Victoria Botuyan / Georges Mer /  |
| PubMed Abstract | RNF168 plays a central role in the DNA damage response (DDR) by ubiquitylating histone H2A at K13 and K15. These modifications direct BRCA1-BARD1 and 53BP1 foci formation in chromatin, essential for ...RNF168 plays a central role in the DNA damage response (DDR) by ubiquitylating histone H2A at K13 and K15. These modifications direct BRCA1-BARD1 and 53BP1 foci formation in chromatin, essential for cell-cycle-dependent DNA double-strand break (DSB) repair pathway selection. The mechanism by which RNF168 catalyzes the targeted accumulation of H2A ubiquitin conjugates to form repair foci around DSBs remains unclear. Here, using cryoelectron microscopy (cryo-EM), nuclear magnetic resonance (NMR) spectroscopy, and functional assays, we provide a molecular description of the reaction cycle and dynamics of RNF168 as it modifies the nucleosome and recognizes its ubiquitylation products. We demonstrate an interaction of a canonical ubiquitin-binding domain within full-length RNF168, which not only engages ubiquitin but also the nucleosome surface, clarifying how such site-specific ubiquitin recognition propels a signal amplification loop. Beyond offering mechanistic insights into a key DDR protein, our study aids in understanding site specificity in both generating and interpreting chromatin ubiquitylation. |
 External links External links |  Mol Cell / Mol Cell /  PubMed:38242129 / PubMed:38242129 /  PubMed Central PubMed Central |
| Methods | EM (single particle) / X-ray diffraction |
| Resolution | 2.049 - 4.0 Å |
| Structure data | EMDB-40604, PDB-8smw: EMDB-40605, PDB-8smx: EMDB-40606, PDB-8smy: EMDB-40607, PDB-8smz: EMDB-40608, PDB-8sn0: EMDB-40609, PDB-8sn1: EMDB-40610, PDB-8sn2: EMDB-40611, PDB-8sn3: EMDB-40612, PDB-8sn4: EMDB-40613, PDB-8sn5: EMDB-40614, PDB-8sn6: EMDB-40615, PDB-8sn7: EMDB-40616, PDB-8sn8: EMDB-40617, PDB-8sn9: EMDB-40618, PDB-8sna: EMDB-41706, PDB-8txv: EMDB-41707, PDB-8txw: EMDB-41708, PDB-8txx: EMDB-41800, PDB-8u13: EMDB-41801, PDB-8u14: EMDB-42446, PDB-8upf:  PDB-8uq8:  PDB-8uq9:  PDB-8uqa:  PDB-8uqb:  PDB-8uqc:  PDB-8uqd:  PDB-8uqe: |
| Chemicals |  ChemComp-ZN:  ChemComp-CL:  ChemComp-GOL:  ChemComp-NA:  ChemComp-HOH: |
| Source |
|
 Keywords Keywords |  STRUCTURAL PROTEIN/DNA/TRANSFERASE / STRUCTURAL PROTEIN/DNA/TRANSFERASE /  Nucleosome core particle / Nucleosome core particle /  chromatin / RNF168 / chromatin / RNF168 /  RING domain / UbcH5c / RING domain / UbcH5c /  DNA repair / DNA double-strand break / DNA repair / DNA double-strand break /  Homologous recombination / Homologous recombination /  53BP1 / 53BP1 /  ubiquitin / ubiquitin /  STRUCTURAL PROTEIN-DNA-TRANSFERASE complex / TRANSFERASE/DNA / STRUCTURAL PROTEIN-DNA-TRANSFERASE complex / TRANSFERASE/DNA /  TRANSFERASE / TRANSFERASE-DNA complex / MIU2-LRM domains / BRCA1-BARD1 / TRANSFERASE / TRANSFERASE-DNA complex / MIU2-LRM domains / BRCA1-BARD1 /  Histone H2A / Histone H2A /  Histone H2B / Histone H2B /  Ubiquitin ligase / Ubiquitin ligase /  Ubiquitin-conjugating enzyme / DNA damage response / DNA double-strand break repair / Ubiquitin-conjugating enzyme / DNA damage response / DNA double-strand break repair /  PROTEIN BINDING / PROTEIN BINDING-Transferase complex PROTEIN BINDING / PROTEIN BINDING-Transferase complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers