-検索条件
-検索結果
検索 (著者・登録者: ai & hs)の結果945件中、1から50件目までを表示しています

EMDB-40815: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies C3V5, V1V3, N611 and base from participant 017

EMDB-40816: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies gp41-N611/FP and base from participant 03

EMDB-40817: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies gp41-N611/FP and base from participant 07

EMDB-40818: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies gp120-GH and base from participant 09

EMDB-40819: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies C3V5, V1V3, gp41-GH/FP and base from participant 11

EMDB-40248: 
CRISPR-Cas type III-D effector complex

EMDB-40250: 
CRISPR-Cas type III-D effector complex bound to a self-target RNA in the pre-cleavage state
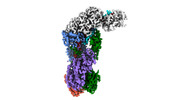
EMDB-40251: 
CRISPR-Cas type III-D effector complex bound to self-target RNA in a post-cleavage state

EMDB-40276: 
CRISPR-Cas type III-D effector complex consensus map

EMDB-40296: 
CRISPR-Cas type III-D effector complex local refinement map

EMDB-40297: 
CRISPR-Cas type III-D effector complex bound to a target RNA local refinement map

EMDB-40298: 
CRISPR-Cas type III-D effector complex bound to a target RNA consensus map

PDB-8s9t: 
CRISPR-Cas type III-D effector complex

PDB-8s9v: 
CRISPR-Cas type III-D effector complex bound to a self-target RNA in the pre-cleavage state

PDB-8s9x: 
CRISPR-Cas type III-D effector complex bound to self-target RNA in a post-cleavage state

EMDB-18334: 
Cryo-EM structure of the inward-facing FLVCR1

EMDB-18335: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1

EMDB-18336: 
Cryo-EM structure of the inward-facing FLVCR2
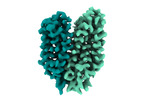
EMDB-18337: 
Cryo-EM structure of the outward-facing FLVCR2

EMDB-18339: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2

EMDB-19009: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1

PDB-8qcs: 
Cryo-EM structure of the inward-facing FLVCR1

PDB-8qct: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1

PDB-8qcx: 
Cryo-EM structure of the inward-facing FLVCR2

PDB-8qcy: 
Cryo-EM structure of the outward-facing FLVCR2

PDB-8qd0: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2

PDB-8r8t: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1

EMDB-35929: 
Cryo-EM structure of Ufd4 in complex with K29/48 triUb

EMDB-35931: 
cryo-EM structures of Ufd4 in complex with Ubc4-Ub

EMDB-36800: 
Potassium transporter KtrAB from Bacillus subtilis in ADP-bound state

EMDB-36801: 
Potassium transporter KtrAB from Bacillus subtilis in ADP-bound state, focused refined on KtrA octamer

EMDB-36802: 
Potassium transporter KtrAB from Bacillus subtilis in ADP-bound state, focused refined on KtrB dimer

EMDB-36803: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of MgCl2

EMDB-36804: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA

EMDB-38477: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA, vertical C2 symmetry axis

EMDB-38478: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA, C1 symmetry

PDB-8k1s: 
Potassium transporter KtrAB from Bacillus subtilis in ADP-bound state

PDB-8k1t: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of MgCl2

PDB-8k1u: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA

PDB-8xmh: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA, vertical C2 symmetry axis

PDB-8xmi: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA, C1 symmetry

EMDB-38330: 
ETB-eGt complex bound to endothelin-1

EMDB-19477: 
Saccharomyces cerevisiae FAS type I

EMDB-19489: 
Tobacco mosaic virus from scanning transmission electron microscopy at CSA=2.0 mrad

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
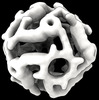
EMDB-42031: 
Computational Designed Nanocage O43_129_+8

EMDB-43318: 
Twistless helix 12 repeat ring design R12B

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage

EMDB-42906: 
Computational Designed Nanocage O43_129
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
