-Search query
-Search result
Showing 1 - 50 of 533 items for (author: moore & a)

EMDB-40815: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies C3V5, V1V3, N611 and base from participant 017
Method: single particle / : Karlinsey D, Ozorowski G, Ward AB

EMDB-40816: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies gp41-N611/FP and base from participant 03
Method: single particle / : Karlinsey D, Ozorowski G, Ward AB

EMDB-40817: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies gp41-N611/FP and base from participant 07
Method: single particle / : Karlinsey D, Ozorowski G, Ward AB

EMDB-40818: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies gp120-GH and base from participant 09
Method: single particle / : Karlinsey D, Ozorowski G, Ward AB

EMDB-40819: 
BG505 SOSIP.664 in complex with wk26 human polyclonal antibodies C3V5, V1V3, gp41-GH/FP and base from participant 11
Method: single particle / : Karlinsey D, Ozorowski G, Ward AB

EMDB-19042: 
MAP7 MTBD (microtubule binding domain) decorated microtubule protofilament
Method: single particle / : Bangera M, Moores CA

EMDB-19043: 
MAP7 MTBD (microtubule binding domain) decorated Taxol stabilized microtubule (13pf) protofilament
Method: single particle / : Bangera M, Moores CA

EMDB-19044: 
MAP7 MTBD (microtubule binding domain) decorated Taxol stabilized microtubule (14pf) protofilament
Method: single particle / : Bangera M, Moores CA

PDB-8rc1: 
MAP7 MTBD (microtubule binding domain) decorated microtubule protofilament
Method: single particle / : Bangera M, Moores CA

EMDB-41888: 
Structure of Apo CXCR4/Gi complex
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41889: 
Structure of CXCL12-bound CXCR4/Gi complex
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41890: 
Structure of AMD3100-bound CXCR4/Gi complex
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41891: 
Structure of REGN7663 Fab-bound CXCR4/Gi complex
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41892: 
Structure of REGN7663-Fab bound CXCR4
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41893: 
Structure of trimeric CXCR4 in complex with REGN7663 Fab
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41894: 
Structure of tetrameric CXCR4 in complex with REGN7663 Fab
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-17557: 
Cryo-EM structure of cortactin-stabilized Arp2/3-complex nucleated actin branches-Local refined map on mother filament
Method: single particle / : Liu T, Moores CA

EMDB-17553: 
Cryo-EM structure of cortactin-stabilized Arp2/3 complex nucleated actin branches-Daughter filament consensus map
Method: single particle / : Liu T, Moores CA

EMDB-17554: 
Cryo-EM structure of cortactin-stabilized Arp2/3 nucleated actin branches-Local refined map on Arp2/3 complex
Method: single particle / : Liu T, Moores CA

EMDB-17555: 
Cryo-EM structure of cortactin-stabilized Arp2/3-complex nucleated actin branches-Local refined map on the daughter filament and cortactin density
Method: single particle / : Liu T, Moores CA

EMDB-17556: 
Cryo-EM structure of cortactin-stabilized Arp2/3-complex nucleated actin branches-Local refined map on capping protein
Method: single particle / : Liu T, Moores CA

EMDB-17558: 
Cryo-EM structure of cortactin stabilized Arp2/3-complex nucleated actin branches
Method: single particle / : Liu T, Moores CA

PDB-8p94: 
Cryo-EM structure of cortactin stabilized Arp2/3-complex nucleated actin branches
Method: single particle / : Liu T, Moores CA

EMDB-42124: 
Cryo-EM structure of human STEAP1 in complex with AMG 509 Fab
Method: single particle / : Li F, Bailis JM

PDB-8ucd: 
Cryo-EM structure of human STEAP1 in complex with AMG 509 Fab
Method: single particle / : Li F, Bailis JM, Zhang H

EMDB-40088: 
HIV-1 Env subtype C CZA97.12 SOSIP.664 in complex with 3BNC117 Fab
Method: single particle / : Ozorowski G, Lee JH, Ward AB

PDB-8gje: 
HIV-1 Env subtype C CZA97.12 SOSIP.664 in complex with 3BNC117 Fab
Method: single particle / : Ozorowski G, Lee JH, Ward AB

EMDB-27703: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
Method: single particle / : May AJ, Manne K, Acharya P

PDB-8dtk: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
Method: single particle / : May AJ, Manne K, Acharya P
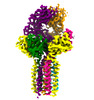
EMDB-28570: 
CryoEM structure of PN45545 TCR-CD3 complex
Method: single particle / : Saotome K, Franklin MC
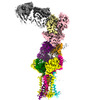
EMDB-28571: 
CryoEM structure of PN45545 TCR-CD3 in complex with HLA-A2 MAGEA4 (230-239)
Method: single particle / : Saotome K, Franklin MC

EMDB-28572: 
CryoEM structure of PN45428 TCR-CD3 in complex with HLA-A2 MAGEA4
Method: single particle / : Saotome K, Franklin MC
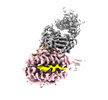
EMDB-28573: 
CryoEM structure of HLA-A2 bound to MAGEA4 (230-239) peptide
Method: single particle / : Saotome K, Franklin MC

EMDB-28574: 
CryoEM structure of HLA-A2 bound to MAGEA8 (232-241) peptide
Method: single particle / : Saotome K, Franklin MC

PDB-8es7: 
CryoEM structure of PN45545 TCR-CD3 complex
Method: single particle / : Saotome K, Franklin MC

PDB-8es8: 
CryoEM structure of PN45545 TCR-CD3 in complex with HLA-A2 MAGEA4 (230-239)
Method: single particle / : Saotome K, Franklin MC

PDB-8es9: 
CryoEM structure of PN45428 TCR-CD3 in complex with HLA-A2 MAGEA4
Method: single particle / : Saotome K, Franklin MC

PDB-8esa: 
CryoEM structure of HLA-A2 bound to MAGEA4 (230-239) peptide
Method: single particle / : Saotome K, Franklin MC

PDB-8esb: 
CryoEM structure of HLA-A2 bound to MAGEA8 (232-241) peptide
Method: single particle / : Saotome K, Franklin MC

PDB-8sbe: 
Structure of the rat vesicular glutamate transporter 2 determined by single-particle Cryo-EM
Method: single particle / : Li F, Finer-Moore J, Eriksen J, Cheng Y, Edwards R, Stroud R

EMDB-27232: 
Subtomogram of Fluad vaccine hemagglutinin spiked nanodisc
Method: electron tomography / : Gallagher JR, Audray AK

EMDB-27233: 
Tomogram of Flublok vaccine hemagglutinin starfish complexes
Method: electron tomography / : Gallagher JR, Audray AK

EMDB-26574: 
KS-AT di-domain of mycobacterial Pks13 with endogenous KS ligand bound
Method: single particle / : Kim SK, Dickinson MS, Finer-Moore JS, Rosenberg OS, Stroud RM

EMDB-27002: 
ACP1-KS-AT domains of mycobacterial Pks13
Method: single particle / : Kim SK, Dickinson MS, Finer-Moore JS, Rosenberg OS, Stroud RM

EMDB-27003: 
KS-AT domains of mycobacterial Pks13 with inward AT conformation
Method: single particle / : Kim SK, Dickinson MS, Finer-Moore JS, Rosenberg OS, Stroud RM

EMDB-27004: 
KS-AT domains of mycobacterial Pks13 with outward AT conformation
Method: single particle / : Kim SK, Dickinson MS, Finer-Moore JS, Rosenberg OS, Stroud RM

EMDB-27005: 
ACP1-KS-AT domains of mycobacterial Pks13
Method: single particle / : Kim SK, Dickinson MS, Finer-Moore JS, Rosenberg OS, Stroud RM

PDB-7uk4: 
KS-AT di-domain of mycobacterial Pks13 with endogenous KS ligand bound
Method: single particle / : Kim SK, Dickinson MS, Finer-Moore JS, Rosenberg OS, Stroud RM

PDB-8cuy: 
ACP1-KS-AT domains of mycobacterial Pks13
Method: single particle / : Kim SK, Dickinson MS, Finer-Moore JS, Rosenberg OS, Stroud RM

PDB-8cuz: 
KS-AT domains of mycobacterial Pks13 with inward AT conformation
Method: single particle / : Kim SK, Dickinson MS, Finer-Moore JS, Rosenberg OS, Stroud RM
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
