-Search query
-Search result
Showing 1 - 50 of 236 items for (author: akashi & s)

EMDB-34823: 
Structure of human SGLT2-MAP17 complex with Phlorizin
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

PDB-8hin: 
Structure of human SGLT2-MAP17 complex with Phlorizin
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

EMDB-34705: 
Structure of human SGLT2-MAP17 complex with Dapagliflozin
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

EMDB-34737: 
Structure of human SGLT2-MAP17 complex with Sotagliflozin
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

PDB-8hez: 
Structure of human SGLT2-MAP17 complex with Dapagliflozin
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

PDB-8hg7: 
Structure of human SGLT2-MAP17 complex with Sotagliflozin
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

EMDB-34673: 
Structure of human SGLT2-MAP17 complex with Canagliflozin
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

PDB-8hdh: 
Structure of human SGLT2-MAP17 complex with Canagliflozin
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

EMDB-34610: 
Structure of human SGLT2-MAP17 complex with TA1887
Method: single particle / : Hiraizumi M, Kishida H, Masahiro I, Nureki O

PDB-8hb0: 
Structure of human SGLT2-MAP17 complex with TA1887
Method: single particle / : Hiraizumi M, Kishida H, Miyaguchi I, Nureki O

EMDB-34530: 
Membrane protein A
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

EMDB-34531: 
Membrane protein B
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

EMDB-35713: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 H225F mutant in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Nakamura S, Yamashita K, Fukuda M, Deisseroth K, Kato HE

PDB-8h86: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

PDB-8h87: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR2 in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

PDB-8iu0: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 H225F mutant in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Nakamura S, Yamashita K, Fukuda M, Deisseroth K, Kato HE

EMDB-35351: 
Cryo-EM structure of CXCL8 bound C-X-C chemokine receptor 1 in complex with Gi heterotrimer
Method: single particle / : Ishimoto N, Park JH, Park SY

PDB-8ic0: 
Cryo-EM structure of CXCL8 bound C-X-C chemokine receptor 1 in complex with Gi heterotrimer
Method: single particle / : Ishimoto N, Park JH, Park SY

EMDB-34588: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 1 (3.2 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34589: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 2 (3.9 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34590: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 4 (3.9 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Sato S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34591: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 1 (4.7 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34592: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 2 (4.8 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34593: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 3
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Sato S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34594: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 4 (4.5 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34595: 
Cryo-EM structure of the CBP catalytic core bound to the H4K12acK16ac nucleosome, class 1
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34596: 
Cryo-EM structure of the CBP catalytic core bound to the H4K12acK16ac nucleosome, class 2
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34597: 
Cryo-EM structure of the CBP catalytic core bound to the H4K12acK16ac nucleosome, class 3
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

PDB-8hag: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 1 (3.2 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

PDB-8hah: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 2 (3.9 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

PDB-8hai: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 1 (4.7 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

PDB-8haj: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 2 (4.8 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

PDB-8hak: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 4 (4.5 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

PDB-8hal: 
Cryo-EM structure of the CBP catalytic core bound to the H4K12acK16ac nucleosome, class 1
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

PDB-8ham: 
Cryo-EM structure of the CBP catalytic core bound to the H4K12acK16ac nucleosome, class 2
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

PDB-8han: 
Cryo-EM structure of the CBP catalytic core bound to the H4K12acK16ac nucleosome, class 3
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T
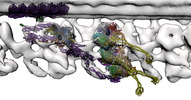
EMDB-16304: 
In situ outer dynein arm from Chlamydomonas reinhardtii in a pre-power stroke state
Method: subtomogram averaging / : Zimmermann NEL, Noga A, Obbineni JM, Ishikawa T
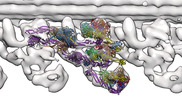
EMDB-16312: 
In situ outer dynein arm from Chlamydomonas reinhardtii in the post-power stroke state
Method: subtomogram averaging / : Zimmermann NEL, Noga A, Obbineni JM, Ishikawa T

PDB-8bwy: 
In situ outer dynein arm from Chlamydomonas reinhardtii in a pre-power stroke state
Method: subtomogram averaging / : Zimmermann NEL, Noga A, Obbineni JM, Ishikawa T

PDB-8bx8: 
In situ outer dynein arm from Chlamydomonas reinhardtii in the post-power stroke state
Method: subtomogram averaging / : Zimmermann NEL, Noga A, Obbineni JM, Ishikawa T

EMDB-16309: 
In situ outer dynein arm from Chlamydomonas reinhardtii in an intermediate state
Method: subtomogram averaging / : Zimmermann NEL, Noga A, Obbineni JM, Ishikawa T

EMDB-16310: 
In situ outer dynein arm from Chlamydomonas reinhardtii in an intermediate
Method: subtomogram averaging / : Zimmermann NEL, Noga A, Obbineni JM, Ishikawa T
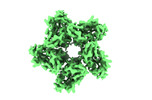
EMDB-35143: 
Cryo-EM structure of the zeaxanthin-bound kin4B8
Method: single particle / : Murakoshi S, Chazan A, Shihoya W, Beja O, Nureki O

PDB-8i2z: 
Cryo-EM structure of the zeaxanthin-bound kin4B8
Method: single particle / : Murakoshi S, Chazan A, Shihoya W, Beja O, Nureki O

EMDB-32094: 
Cryo-EM structure of H2A-ubiquitinated nucleosome
Method: single particle / : Ohtomo H, Ito S, McKenzie NJ, Uckelmann M, Wakamori M, Ehara H, Tsunaka Y, Shibata M, Sekine S, Umehara T, Davidovich C, Koseki H, Nishimura Y

EMDB-16022: 
Amyloid-beta 42 filaments extracted from the human brain with Arctic mutation (E22G) of Alzheimer's disease | ABeta42
Method: helical / : Yang Y, Zhang WJ, Murzin AG, Schweighauser M, Huang M, Lovestam SKA, Peak-Chew SY, Macdonald J, Lavenir I, Ghetti B, Graff C, Kumar A, Nordberg A, Goedert M, Scheres SHW

EMDB-16023: 
Amyloid-beta tetrameric filaments with the Arctic mutation (E22G) from Alzheimer's disease brains | ABeta40
Method: helical / : Yang Y, Zhang WJ, Murzin AG, Schweighauser M, Huang M, Lovestam SKA, Peak-Chew SY, Macdonald J, Lavenir I, Ghetti B, Graff C, Kumar A, Nordber A, Goedert M, Scheres SHW

EMDB-16027: 
Murine amyloid-beta filaments with the Arctic mutation (E22G) from APP(NL-G-F) mouse brains | ABeta
Method: helical / : Yang Y, Zhang WJ, Murzin AG, Schweighauser M, Huang M, Lovestam SKA, Peak-Chew SY, Macdonald J, Lavenir I, Ghetti B, Graff C, Kumar A, Nordber A, Goedert M, Scheres SHW

PDB-8bfz: 
Amyloid-beta 42 filaments extracted from the human brain with Arctic mutation (E22G) of Alzheimer's disease | ABeta42
Method: helical / : Yang Y, Zhang WJ, Murzin AG, Schweighauser M, Huang M, Lovestam SKA, Peak-Chew SY, Macdonald J, Lavenir I, Ghetti B, Graff C, Kumar A, Nordberg A, Goedert M, Scheres SHW

PDB-8bg0: 
Amyloid-beta tetrameric filaments with the Arctic mutation (E22G) from Alzheimer's disease brains | ABeta40
Method: helical / : Yang Y, Zhang WJ, Murzin AG, Schweighauser M, Huang M, Lovestam SKA, Peak-Chew SY, Macdonald J, Lavenir I, Ghetti B, Graff C, Kumar A, Nordber A, Goedert M, Scheres SHW
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
