+ Open data
Open data
- Basic information
Basic information
| Entry | 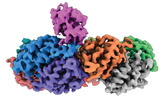 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
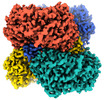
| Title | Cryo-EM structure of the human PRDX4-ErP46 complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Peroxiredoxin-4 / Peroxiredoxin-4 /  OXIDOREDUCTASE OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology information Oxidoreductases; Acting on a sulfur group of donors; With a disulfide as acceptor / cellular response to stress / Oxidoreductases; Acting on a sulfur group of donors; With a disulfide as acceptor / cellular response to stress /  protein disulfide-isomerase / negative regulation of male germ cell proliferation / I-kappaB phosphorylation / thioredoxin-dependent peroxiredoxin / thioredoxin peroxidase activity / molecular sequestering activity / protein maturation by protein folding / Lysosome Vesicle Biogenesis ... protein disulfide-isomerase / negative regulation of male germ cell proliferation / I-kappaB phosphorylation / thioredoxin-dependent peroxiredoxin / thioredoxin peroxidase activity / molecular sequestering activity / protein maturation by protein folding / Lysosome Vesicle Biogenesis ... Oxidoreductases; Acting on a sulfur group of donors; With a disulfide as acceptor / cellular response to stress / Oxidoreductases; Acting on a sulfur group of donors; With a disulfide as acceptor / cellular response to stress /  protein disulfide-isomerase / negative regulation of male germ cell proliferation / I-kappaB phosphorylation / thioredoxin-dependent peroxiredoxin / thioredoxin peroxidase activity / molecular sequestering activity / protein maturation by protein folding / Lysosome Vesicle Biogenesis / Golgi Associated Vesicle Biogenesis / protein disulfide-isomerase / negative regulation of male germ cell proliferation / I-kappaB phosphorylation / thioredoxin-dependent peroxiredoxin / thioredoxin peroxidase activity / molecular sequestering activity / protein maturation by protein folding / Lysosome Vesicle Biogenesis / Golgi Associated Vesicle Biogenesis /  protein disulfide isomerase activity / protein disulfide isomerase activity /  protein-disulfide reductase activity / extracellular matrix organization / lysosomal lumen / cell redox homeostasis / hydrogen peroxide catabolic process / male gonad development / azurophil granule lumen / protein-disulfide reductase activity / extracellular matrix organization / lysosomal lumen / cell redox homeostasis / hydrogen peroxide catabolic process / male gonad development / azurophil granule lumen /  protein folding / protein folding /  spermatogenesis / secretory granule lumen / response to oxidative stress / ficolin-1-rich granule lumen / molecular adaptor activity / spermatogenesis / secretory granule lumen / response to oxidative stress / ficolin-1-rich granule lumen / molecular adaptor activity /  endoplasmic reticulum lumen / Neutrophil degranulation / negative regulation of apoptotic process / endoplasmic reticulum lumen / Neutrophil degranulation / negative regulation of apoptotic process /  endoplasmic reticulum / extracellular exosome / extracellular region / identical protein binding / endoplasmic reticulum / extracellular exosome / extracellular region / identical protein binding /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.47 Å cryo EM / Resolution: 3.47 Å | |||||||||
 Authors Authors | Su CC / Lyu M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2023 Journal: Cell Rep / Year: 2023Title: High-resolution structural-omics of human liver enzymes. Authors: Chih-Chia Su / Meinan Lyu / Zhemin Zhang / Masaru Miyagi / Wei Huang / Derek J Taylor / Edward W Yu /  Abstract: We applied raw human liver microsome lysate to a holey carbon grid and used cryo-electron microscopy (cryo-EM) to define its composition. From this sample we identified and simultaneously determined ...We applied raw human liver microsome lysate to a holey carbon grid and used cryo-electron microscopy (cryo-EM) to define its composition. From this sample we identified and simultaneously determined high-resolution structural information for ten unique human liver enzymes involved in diverse cellular processes. Notably, we determined the structure of the endoplasmic bifunctional protein H6PD, where the N- and C-terminal domains independently possess glucose-6-phosphate dehydrogenase and 6-phosphogluconolactonase enzymatic activity, respectively. We also obtained the structure of heterodimeric human GANAB, an ER glycoprotein quality-control machinery that contains a catalytic α subunit and a noncatalytic β subunit. In addition, we observed a decameric peroxidase, PRDX4, which directly contacts a disulfide isomerase-related protein, ERp46. Structural data suggest that several glycosylations, bound endogenous compounds, and ions associate with these human liver enzymes. These results highlight the importance of cryo-EM in facilitating the elucidation of human organ proteomics at the atomic level. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28217.map.gz emd_28217.map.gz | 97.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28217-v30.xml emd-28217-v30.xml emd-28217.xml emd-28217.xml | 16.9 KB 16.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28217_fsc.xml emd_28217_fsc.xml emd_28217_fsc_2.xml emd_28217_fsc_2.xml | 9.9 KB 13.8 KB | Display Display |  FSC data file FSC data file |
| Images |  emd_28217.png emd_28217.png | 103.2 KB | ||
| Masks |  emd_28217_msk_1.map emd_28217_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-28217.cif.gz emd-28217.cif.gz | 5.5 KB | ||
| Others |  emd_28217_additional_1.map.gz emd_28217_additional_1.map.gz emd_28217_half_map_1.map.gz emd_28217_half_map_1.map.gz emd_28217_half_map_2.map.gz emd_28217_half_map_2.map.gz | 51.8 MB 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28217 http://ftp.pdbj.org/pub/emdb/structures/EMD-28217 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28217 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28217 | HTTPS FTP |
-Related structure data
| Related structure data |  8ekyMC  7uzmC  8ekwC  8em2C  8emrC  8emsC  8emtC  8eneC  8eojC  8eorC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28217.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28217.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28217_msk_1.map emd_28217_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened map
| File | emd_28217_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_28217_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_28217_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : H6PD
| Entire | Name: H6PD |
|---|---|
| Components |
|
-Supramolecule #1: H6PD
| Supramolecule | Name: H6PD / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Peroxiredoxin-4
| Macromolecule | Name: Peroxiredoxin-4 / type: protein_or_peptide / ID: 1 / Number of copies: 10 / Enantiomer: LEVO / EC number: thioredoxin-dependent peroxiredoxin |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 30.578873 KDa |
| Sequence | String: MEALPLLAAT TPDHGRHRRL LLLPLLLFLL PAGAVQGWET EERPRTREEE CHFYAGGQVY PGEASRVSVA DHSLHLSKAK ISKPAPYWE GTAVIDGEFK ELKLTDYRGK YLVFFFYPLD FTFVCPTEII AFGDRLEEFR SINTEVVACS VDSQFTHLAW I NTPRRQGG ...String: MEALPLLAAT TPDHGRHRRL LLLPLLLFLL PAGAVQGWET EERPRTREEE CHFYAGGQVY PGEASRVSVA DHSLHLSKAK ISKPAPYWE GTAVIDGEFK ELKLTDYRGK YLVFFFYPLD FTFVCPTEII AFGDRLEEFR SINTEVVACS VDSQFTHLAW I NTPRRQGG LGPIRIPLLS DLTHQISKDY GVYLEDSGHT LRGLFIIDDK GILRQITLND LPVGRSVDET LRLVQAFQYT DK HGEVCPA GWKPGSETII PDPAGKLKYF DKLN UniProtKB:  Peroxiredoxin-4 Peroxiredoxin-4 |
-Macromolecule #2: Thioredoxin domain-containing protein 5
| Macromolecule | Name: Thioredoxin domain-containing protein 5 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO EC number:  Oxidoreductases; Acting on a sulfur group of donors; With a disulfide as acceptor Oxidoreductases; Acting on a sulfur group of donors; With a disulfide as acceptor |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.687727 KDa |
| Sequence | String: MPARPGRLLP LLARPAALTA LLLLLLGHGG GGRWGARAQE AAAAAADGPP AADGEDGQDP HSKHLYTADM FTHGIQSAAH FVMFFAPWC GHCQRLQPTW NDLGDKYNSM EDAKVYVAKV DCTAHSDVCS AQGVRGYPTL KLFKPGQEAV KYQGPRDFQT L ENWMLQTL ...String: MPARPGRLLP LLARPAALTA LLLLLLGHGG GGRWGARAQE AAAAAADGPP AADGEDGQDP HSKHLYTADM FTHGIQSAAH FVMFFAPWC GHCQRLQPTW NDLGDKYNSM EDAKVYVAKV DCTAHSDVCS AQGVRGYPTL KLFKPGQEAV KYQGPRDFQT L ENWMLQTL NEEPVTPEPE VEPPSAPELK QGLYELSASN FELHVAQGDH FIKFFAPWCG HCKALAPTWE QLALGLEHSE TV KIGKVDC TQHYELCSGN QVRGYPTLLW FRDGKKVDQY KGKRDLESLR EYVESQLQRT ETGATETVTP SEAPVLAAEP EAD KGTVLA LTENNFDDTI AEGITFIKFY APWCGHCKTL APTWEELSKK EFPGLAGVKI AEVDCTAERN ICSKYSVRGY PTLL LFRGG KKVSEHSGGR DLDSLHRFVL SQAKDEL UniProtKB: Thioredoxin domain-containing protein 5 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | This is from a heterogeneous and impure protein sample. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 29.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8eky: |
 Movie
Movie Controller
Controller



















 Z
Z Y
Y X
X


































