+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of double homo-hexameric AAA+ ATPase RuvB motors | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information Holliday junction resolvase complex / four-way junction helicase activity / four-way junction DNA binding / DNA recombination / Holliday junction resolvase complex / four-way junction helicase activity / four-way junction DNA binding / DNA recombination /  DNA helicase / DNA helicase /  DNA repair / DNA repair /  ATP hydrolysis activity / ATP hydrolysis activity /  ATP binding / ATP binding /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |    Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) | |||||||||
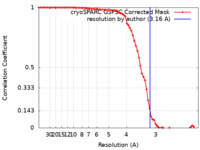
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.16 Å cryo EM / Resolution: 3.16 Å | |||||||||
 Authors Authors | Shen ZF / Rish AD / Fu TM | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of double homo-hexameric AAA+ ATPase RuvB motor binding with DNA substrate Authors: Rish AD / Shen ZF / Fu TM | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28107.map.gz emd_28107.map.gz | 36.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28107-v30.xml emd-28107-v30.xml emd-28107.xml emd-28107.xml | 15.7 KB 15.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28107_fsc.xml emd_28107_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_28107.png emd_28107.png | 120.6 KB | ||
| Masks |  emd_28107_msk_1.map emd_28107_msk_1.map | 38.4 MB |  Mask map Mask map | |
| Others |  emd_28107_half_map_1.map.gz emd_28107_half_map_1.map.gz emd_28107_half_map_2.map.gz emd_28107_half_map_2.map.gz | 35.5 MB 35.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28107 http://ftp.pdbj.org/pub/emdb/structures/EMD-28107 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28107 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28107 | HTTPS FTP |
-Related structure data
| Related structure data |  8efyMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28107.map.gz / Format: CCP4 / Size: 38.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28107.map.gz / Format: CCP4 / Size: 38.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28107_msk_1.map emd_28107_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_28107_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_28107_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Double homo-hexameric AAA+ ATPase RuvB motors
| Entire | Name: Double homo-hexameric AAA+ ATPase RuvB motors |
|---|---|
| Components |
|
-Supramolecule #1: Double homo-hexameric AAA+ ATPase RuvB motors
| Supramolecule | Name: Double homo-hexameric AAA+ ATPase RuvB motors / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:    Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) |
-Macromolecule #1: Holliday junction ATP-dependent DNA helicase RuvB
| Macromolecule | Name: Holliday junction ATP-dependent DNA helicase RuvB / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO / EC number:  DNA helicase DNA helicase |
|---|---|
| Source (natural) | Organism:    Thermus thermophilus HB8 (bacteria) / Strain: ATCC 27634 / DSM 579 / HB8 Thermus thermophilus HB8 (bacteria) / Strain: ATCC 27634 / DSM 579 / HB8 |
| Molecular weight | Theoretical: 36.024688 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MEDLALRPKT LDEYIGQERL KQKLRVYLEA AKARKEPLEH LLLFGPPGLG KTTLAHVIAH ELGVNLRVTS GPAIEKPGDL AAILANSLE EGDILFIDEI HRLSRQAEEH LYPAMEDFVM DIVIGQGPAA RTIRLELPRF TLIGATTRPG LITAPLLSRF G IVEHLEYY ...String: MEDLALRPKT LDEYIGQERL KQKLRVYLEA AKARKEPLEH LLLFGPPGLG KTTLAHVIAH ELGVNLRVTS GPAIEKPGDL AAILANSLE EGDILFIDEI HRLSRQAEEH LYPAMEDFVM DIVIGQGPAA RTIRLELPRF TLIGATTRPG LITAPLLSRF G IVEHLEYY TPEELAQGVM RDARLLGVRI TEEAALEIGR RSRGTMRVAK RLFRRVRDFA QVAGEEVITR ERALEALAAL GL DELGLEK RDREILEVLI LRFGGGPVGL ATLATALSED PGTLEEVHEP YLIRQGLLKR TPRGRVATEL AYRHLGYPPP VGP LLEP |
-Macromolecule #2: ssDNA
| Macromolecule | Name: ssDNA / type: dna / ID: 2 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:    Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) |
| Molecular weight | Theoretical: 15.146732 KDa |
| Sequence | String: (DA)(DG)(DA)(DA)(DT)(DC)(DT)(DG)(DC)(DC) (DG)(DA)(DG)(DA)(DG)(DA)(DC)(DC)(DG)(DA) (DG)(DC)(DA)(DG)(DA)(DA)(DT)(DT)(DC) (DT)(DA)(DT)(DG)(DT)(DG)(DT)(DT)(DT)(DA) (DC) (DC)(DA)(DA)(DG)(DC)(DG)(DC)(DT) (DG) |
-Macromolecule #3: ssDNA
| Macromolecule | Name: ssDNA / type: dna / ID: 3 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:    Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) |
| Molecular weight | Theoretical: 15.69107 KDa |
| Sequence | String: (DC)(DA)(DG)(DC)(DG)(DC)(DT)(DT)(DG)(DG) (DT)(DA)(DA)(DA)(DC)(DA)(DC)(DA)(DT)(DA) (DG)(DA)(DA)(DT)(DT)(DC)(DT)(DG)(DC) (DT)(DC)(DT)(DC)(DG)(DG)(DT)(DC)(DT)(DG) (DA) (DG)(DC)(DC)(DG)(DT)(DC)(DT)(DA) (DA)(DG)(DA) |
-Macromolecule #4: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 6 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 8 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER / type: ligand / ID: 6 / Number of copies: 6 / Formula: AGS |
|---|---|
| Molecular weight | Theoretical: 523.247 Da |
| Chemical component information |  ChemComp-AGS: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X