[English] 日本語
 Yorodumi
Yorodumi- EMDB-13786: Cryo-EM structure of S.cerevisiae condensin Ycg1-Brn1-DNA complex -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
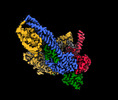
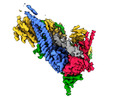
| Title | Cryo-EM structure of S.cerevisiae condensin Ycg1-Brn1-DNA complex | ||||||||||||
 Map data Map data | cropped box=162pix | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of meiotic DNA double-strand break formation / meiotic chromosome condensation / Condensation of Prometaphase Chromosomes / tRNA gene clustering / meiotic chromosome separation /  condensin complex / rDNA chromatin condensation / condensin complex / rDNA chromatin condensation /  synaptonemal complex assembly / mitotic chromosome condensation / chromosome condensation ...negative regulation of meiotic DNA double-strand break formation / meiotic chromosome condensation / Condensation of Prometaphase Chromosomes / tRNA gene clustering / meiotic chromosome separation / synaptonemal complex assembly / mitotic chromosome condensation / chromosome condensation ...negative regulation of meiotic DNA double-strand break formation / meiotic chromosome condensation / Condensation of Prometaphase Chromosomes / tRNA gene clustering / meiotic chromosome separation /  condensin complex / rDNA chromatin condensation / condensin complex / rDNA chromatin condensation /  synaptonemal complex assembly / mitotic chromosome condensation / chromosome condensation / mitotic sister chromatid segregation / condensed chromosome / synaptonemal complex assembly / mitotic chromosome condensation / chromosome condensation / mitotic sister chromatid segregation / condensed chromosome /  cell division / cell division /  chromatin binding / chromatin binding /  nucleus / nucleus /  cytoplasm cytoplasmSimilarity search - Function | ||||||||||||
| Biological species |   Saccharomyces cerevisiae S288C (yeast) / synthetic construct (others) Saccharomyces cerevisiae S288C (yeast) / synthetic construct (others) | ||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.2 Å cryo EM / Resolution: 3.2 Å | ||||||||||||
 Authors Authors | Lee B-G / Rhodes J / Lowe J | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Clamping of DNA shuts the condensin neck gate. Authors: Byung-Gil Lee / James Rhodes / Jan Löwe /  Abstract: SignificanceDNA needs to be compacted to fit into nuclei and during cell division, when dense chromatids are formed for their mechanical segregation, a process that depends on the protein complex ...SignificanceDNA needs to be compacted to fit into nuclei and during cell division, when dense chromatids are formed for their mechanical segregation, a process that depends on the protein complex condensin. It forms and enlarges loops in DNA through loop extrusion. Our work resolves the atomic structure of a DNA-bound state of condensin in which ATP has not been hydrolyzed. The DNA is clamped within a compartment that has been reported previously in other structural maintenance of chromosomes (SMC) complexes, including Rad50, cohesin, and MukBEF. With the caveat of important differences, it means that all SMC complexes cycle through at least some similar states and undergo similar conformational changes in their head modules, while hydrolyzing ATP and translocating DNA. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13786.map.gz emd_13786.map.gz | 15.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13786-v30.xml emd-13786-v30.xml emd-13786.xml emd-13786.xml | 21 KB 21 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13786.png emd_13786.png | 86.7 KB | ||
| Others |  emd_13786_half_map_1.map.gz emd_13786_half_map_1.map.gz emd_13786_half_map_2.map.gz emd_13786_half_map_2.map.gz | 15.1 MB 15.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13786 http://ftp.pdbj.org/pub/emdb/structures/EMD-13786 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13786 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13786 | HTTPS FTP |
-Related structure data
| Related structure data |  7q2zMC  7q2xC  7q2yC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13786.map.gz / Format: CCP4 / Size: 16.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13786.map.gz / Format: CCP4 / Size: 16.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cropped box=162pix | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: cropped box=162pix
| File | emd_13786_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cropped box=162pix | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: cropped box=162pix
| File | emd_13786_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cropped box=162pix | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : complex of condensin ycg1 and DNA
| Entire | Name: complex of condensin ycg1 and DNA |
|---|---|
| Components |
|
-Supramolecule #1: complex of condensin ycg1 and DNA
| Supramolecule | Name: complex of condensin ycg1 and DNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: ycg1-DNA structure from the dataset of complex of condensin pentamer (smc2, smc4, brn1, ycs4, ycg1), DNA, ADP-BeF3 |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae S288C (yeast) Saccharomyces cerevisiae S288C (yeast) |
| Recombinant expression | Organism:   Saccharomyces cerevisiae W303 (yeast) Saccharomyces cerevisiae W303 (yeast) |
| Molecular weight | Experimental: 120 kDa/nm |
-Macromolecule #1: Condensin complex subunit 3
| Macromolecule | Name: Condensin complex subunit 3 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae S288C (yeast) / Strain: ATCC 204508 / S288c Saccharomyces cerevisiae S288C (yeast) / Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 117.981 KDa |
| Recombinant expression | Organism:   Saccharomyces cerevisiae W303 (yeast) Saccharomyces cerevisiae W303 (yeast) |
| Sequence | String: MQDPDGIDIN TKIFNSVAEV FQKAQGSYAG HRKHIAVLKK IQSKAVEQGY EDAFNFWFDK LVTKILPLKK NEIIGDRIVK LVAAFIASL ERELILAKKQ NYKLTNDEEG IFSRFVDQFI RHVLRGVESP DKNVRFRVLQ LLAVIMDNIG EIDESLFNLL I LSLNKRIY ...String: MQDPDGIDIN TKIFNSVAEV FQKAQGSYAG HRKHIAVLKK IQSKAVEQGY EDAFNFWFDK LVTKILPLKK NEIIGDRIVK LVAAFIASL ERELILAKKQ NYKLTNDEEG IFSRFVDQFI RHVLRGVESP DKNVRFRVLQ LLAVIMDNIG EIDESLFNLL I LSLNKRIY DREPTVRIQA VFCLTKFQDE EQTEHLTELS DNEENFEATR TLVASIQNDP SAEVRRAAML NLINDNNTRP YI LERARDV NIVNRRLVYS RILKSMGRKC FDDIEPHIFD QLIEWGLEDR ELSVRNACKR LIAHDWLNAL DGDLIELLEK LDV SRSSVC VKAIEALFQS RPDILSKIKF PESIWKDFTV EIAFLFRAIY LYCLDNNITE MLEENFPEAS KLSEHLNHYI LLRY HHNDI SNDSQSHFDY NTLEFIIEQL SIAAERYDYS DEVGRRSMLT VVRNMLALTT LSEPLIKIGI RVMKSLSINE KDFVT MAIE IINDIRDDDI EKQEQEEKIK SKKINRRNET SVDEEDENGT HNDEVNEDEE DDNISSFHSA VENLVQGNGN VSESDI INN LPPEKEASSA TIVLCLTRSS YMLELVNTPL TENILIASLM DTLITPAVRN TAPNIRELGV KNLGLCCLLD VKLAIDN MY ILGMCVSKGN ASLKYIALQV IVDIFSVHGN TVVDGEGKVD SISLHKIFYK VLKNNGLPEC QVIAAEGLCK LFLADVFT D DDLFETLVLS YFSPINSSNE ALVQAFAFCI PVYCFSHPAH QQRMSRTAAD ILLRLCVLWD DLQSSVIPEV DREAMLKPN IIFQQLLFWT DPRNLVNQTG STKKDTVQLT FLIDVLKIYA QIEKKEIKKM IITNINAIFL SSEQDYSTLK ELLEYSDDIA ENDNLDNVS KNALDKLRNN LNSLIEEINE RSETQTKDEN NTANDQYSSI LGNSFNKSSN DTIEHAADIT DGNNTELTKT T VNISAVDN TTEQSNSRKR TRSEAEQIDT SKNLENMSIQ DTSTVAKNVS FVLPDEKSDA MSIDEEDKDS ESFSEVC |
-Macromolecule #2: Condensin complex subunit 2
| Macromolecule | Name: Condensin complex subunit 2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae S288C (yeast) / Strain: ATCC 204508 / S288c Saccharomyces cerevisiae S288C (yeast) / Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 86.323297 KDa |
| Recombinant expression | Organism:   Saccharomyces cerevisiae W303 (yeast) Saccharomyces cerevisiae W303 (yeast) |
| Sequence | String: MTTQLRYENN DDDERVEYNL FTNRSTMMAN FEEWIKMATD NKINSRNSWN FALIDYFYDL DVLKDGENNI NFQKASATLD GCIKIYSSR VDSVTTETGK LLSGLAQRKT NGASNGDDSN GGNGEGLGGD SDEANIEIDP LTGMPISNDP DVNNTRRRVY N RVLETTLV ...String: MTTQLRYENN DDDERVEYNL FTNRSTMMAN FEEWIKMATD NKINSRNSWN FALIDYFYDL DVLKDGENNI NFQKASATLD GCIKIYSSR VDSVTTETGK LLSGLAQRKT NGASNGDDSN GGNGEGLGGD SDEANIEIDP LTGMPISNDP DVNNTRRRVY N RVLETTLV EFETIKMKEL DQELIIDPLF KKALVDFDEG GAKSLLLNTL NIDNTARVIF DASIKDTQNV GQGKLQRKEE EL IERDSLV DDENEPSQSL ISTRNDSTVN DSVISAPSME DEILSLGMDF IKFDQIAVCE ISGSIEQLRN VVEDINQAKD FIE NVNNRF DNFLTEEELQ AAVPDNAEDD SDGFDMGMQQ ELCYPDENHD NTSHDEQDDD NVNSTTGSIF EKDLMAYFDE NLNR NWRGR EHWKVRNFKK ANLVNKESDL LEETRTTIGD TTDKNTTDDK SMDTKKKHKQ KKVLEIDFFK TDDSFEDKVF ASKGR TKID MPIKNRKNDT HYLLPDDFHF STDRITRLFI KPGQKMSLFS HRKHTRGDVS SGLFEKSTVS ANHSNNDIPT IADEHF WAD NYERKEQEEK EKEQSKEVGD VVGGALDNPF EDDMDGVDFN QAFEGTDDNE EASVKLDLQD DEDHKFPIRE NKVTYSR VS KKVDVRRLKK NVWRSINNLI QEHDSRKNRE QSSNDSETHT EDESTKELKF SDIIQGISKM YSDDTLKDIS TSFCFICL L HLANEHGLQI THTENYNDLI VNYEDLATTQ AAS |
-Macromolecule #3: DNA
| Macromolecule | Name: DNA / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 6.951477 KDa |
| Sequence | String: (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT) |
-Macromolecule #4: DNA
| Macromolecule | Name: DNA / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 7.1588 KDa |
| Sequence | String: (DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA) (DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA) (DA)(DA)(DA) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm Bright-field microscopy / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Software - Name: CTFFIND (ver. 4) |
|---|---|
| Initial angle assignment | Type: ANGULAR RECONSTITUTION / Software: (Name: cryoSPARC, RELION) |
| Final angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 3.2 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 91024 |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X



















