+Search query
-Structure paper
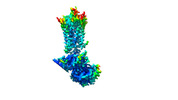
| Title | GPCR activation and GRK2 assembly by a biased intracellular agonist. |
|---|---|
| Journal, issue, pages | Nature, Vol. 620, Issue 7974, Page 676-681, Year 2023 |
| Publish date | Aug 2, 2023 |
 Authors Authors | Jia Duan / Heng Liu / Fenghui Zhao / Qingning Yuan / Yujie Ji / Xiaoqing Cai / Xinheng He / Xinzhu Li / Junrui Li / Kai Wu / Tianyu Gao / Shengnan Zhu / Shi Lin / Ming-Wei Wang / Xi Cheng / Wanchao Yin / Yi Jiang / Dehua Yang / H Eric Xu /  |
| PubMed Abstract | Phosphorylation of G-protein-coupled receptors (GPCRs) by GPCR kinases (GRKs) desensitizes G-protein signalling and promotes arrestin signalling, which is also modulated by biased ligands. The ...Phosphorylation of G-protein-coupled receptors (GPCRs) by GPCR kinases (GRKs) desensitizes G-protein signalling and promotes arrestin signalling, which is also modulated by biased ligands. The molecular assembly of GRKs on GPCRs and the basis of GRK-mediated biased signalling remain largely unknown owing to the weak GPCR-GRK interactions. Here we report the complex structure of neurotensin receptor 1 (NTSR1) bound to GRK2, Gα and the arrestin-biased ligand SBI-553. The density map reveals the arrangement of the intact GRK2 with the receptor, with the N-terminal helix of GRK2 docking into the open cytoplasmic pocket formed by the outward movement of the receptor transmembrane helix 6, analogous to the binding of the G protein to the receptor. SBI-553 binds at the interface between GRK2 and NTSR1 to enhance GRK2 binding. The binding mode of SBI-553 is compatible with arrestin binding but clashes with the binding of Gα protein, thus providing a mechanism for its arrestin-biased signalling capability. In sum, our structure provides a rational model for understanding the details of GPCR-GRK interactions and GRK2-mediated biased signalling. |
 External links External links |  Nature / Nature /  PubMed:37532940 PubMed:37532940 |
| Methods | EM (single particle) |
| Resolution | 2.81 - 3.07 Å |
| Structure data | EMDB-36474, PDB-8jpb: EMDB-36475, PDB-8jpc: EMDB-36476, PDB-8jpd: EMDB-36477, PDB-8jpe: EMDB-36478, PDB-8jpf: |
| Chemicals |  ChemComp-STU: 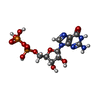 ChemComp-GDP:  ChemComp-MG: 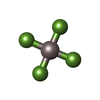 ChemComp-ALF:  ChemComp-SRW: |
| Source |
|
 Keywords Keywords |  SIGNALING PROTEIN / Biased signaling SIGNALING PROTEIN / Biased signaling |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers















