[English] 日本語
 Yorodumi
Yorodumi- PDB-8ki3: Structure of the human ATP synthase bound to bedaquiline (composite) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8ki3 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human ATP synthase bound to bedaquiline (composite) | ||||||||||||
 Components Components |
| ||||||||||||
 Keywords Keywords |  MEMBRANE PROTEIN / MEMBRANE PROTEIN /  ATP synthase / ATP synthase /  Human / Human /  cryo-EM cryo-EM | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationmitochondrial proton-transporting ATP synthase complex binding / regulation of ATP metabolic process / negative regulation of cell adhesion involved in substrate-bound cell migration / regulation of protein targeting to mitochondrion / Formation of ATP by chemiosmotic coupling / Cristae formation / positive regulation of proteolysis involved in protein catabolic process / positive regulation of autophagy of mitochondrion in response to mitochondrial depolarization / ATP biosynthetic process /  angiostatin binding ...mitochondrial proton-transporting ATP synthase complex binding / regulation of ATP metabolic process / negative regulation of cell adhesion involved in substrate-bound cell migration / regulation of protein targeting to mitochondrion / Formation of ATP by chemiosmotic coupling / Cristae formation / positive regulation of proteolysis involved in protein catabolic process / positive regulation of autophagy of mitochondrion in response to mitochondrial depolarization / ATP biosynthetic process / angiostatin binding ...mitochondrial proton-transporting ATP synthase complex binding / regulation of ATP metabolic process / negative regulation of cell adhesion involved in substrate-bound cell migration / regulation of protein targeting to mitochondrion / Formation of ATP by chemiosmotic coupling / Cristae formation / positive regulation of proteolysis involved in protein catabolic process / positive regulation of autophagy of mitochondrion in response to mitochondrial depolarization / ATP biosynthetic process /  angiostatin binding / ATPase inhibitor activity / mitochondrial depolarization / positive regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway / angiostatin binding / ATPase inhibitor activity / mitochondrial depolarization / positive regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway /  Mitochondrial protein import / negative regulation of ATP-dependent activity / mitochondrial proton-transporting ATP synthase complex assembly / negative regulation of hydrolase activity / mitochondrial proton-transporting ATP synthase, catalytic core / mitochondrial proton-transporting ATP synthase, stator stalk / Mitochondrial protein import / negative regulation of ATP-dependent activity / mitochondrial proton-transporting ATP synthase complex assembly / negative regulation of hydrolase activity / mitochondrial proton-transporting ATP synthase, catalytic core / mitochondrial proton-transporting ATP synthase, stator stalk /  enzyme inhibitor activity / proton-transporting ATP synthase complex / cellular response to interleukin-7 / enzyme inhibitor activity / proton-transporting ATP synthase complex / cellular response to interleukin-7 /  oxidative phosphorylation / mitochondrial proton-transporting ATP synthase complex, coupling factor F(o) / response to muscle activity / response to copper ion / heme biosynthetic process / mitochondrial proton-transporting ATP synthase complex / mitochondrial proton-transporting ATP synthase complex, catalytic sector F(1) / mitochondrial nucleoid / proton motive force-driven mitochondrial ATP synthesis / negative regulation of endothelial cell proliferation / proton-transporting ATP synthase complex, coupling factor F(o) / proton motive force-driven ATP synthesis / proton transmembrane transporter activity / cellular response to nitric oxide / response to hyperoxia / proton-transporting ATP synthase complex, catalytic core F(1) / positive regulation of blood vessel endothelial cell migration / MHC class I protein binding / oxidative phosphorylation / mitochondrial proton-transporting ATP synthase complex, coupling factor F(o) / response to muscle activity / response to copper ion / heme biosynthetic process / mitochondrial proton-transporting ATP synthase complex / mitochondrial proton-transporting ATP synthase complex, catalytic sector F(1) / mitochondrial nucleoid / proton motive force-driven mitochondrial ATP synthesis / negative regulation of endothelial cell proliferation / proton-transporting ATP synthase complex, coupling factor F(o) / proton motive force-driven ATP synthesis / proton transmembrane transporter activity / cellular response to nitric oxide / response to hyperoxia / proton-transporting ATP synthase complex, catalytic core F(1) / positive regulation of blood vessel endothelial cell migration / MHC class I protein binding /  aerobic respiration / aerobic respiration /  H+-transporting two-sector ATPase / substantia nigra development / proton-transporting ATPase activity, rotational mechanism / reactive oxygen species metabolic process / proton transmembrane transport / proton-transporting ATP synthase activity, rotational mechanism / cellular response to dexamethasone stimulus / H+-transporting two-sector ATPase / substantia nigra development / proton-transporting ATPase activity, rotational mechanism / reactive oxygen species metabolic process / proton transmembrane transport / proton-transporting ATP synthase activity, rotational mechanism / cellular response to dexamethasone stimulus /  erythrocyte differentiation / generation of precursor metabolites and energy / erythrocyte differentiation / generation of precursor metabolites and energy /  ADP binding / ADP binding /  regulation of intracellular pH / regulation of intracellular pH /  mitochondrial membrane / Transcriptional activation of mitochondrial biogenesis / lipid metabolic process / osteoblast differentiation / mitochondrial membrane / Transcriptional activation of mitochondrial biogenesis / lipid metabolic process / osteoblast differentiation /  ATPase binding / ATPase binding /  angiogenesis / response to ethanol / angiogenesis / response to ethanol /  nuclear membrane / nuclear membrane /  mitochondrial inner membrane / mitochondrial inner membrane /  protease binding / protease binding /  calmodulin binding / calmodulin binding /  hydrolase activity / hydrolase activity /  mitochondrial matrix / mitochondrial matrix /  membrane raft / membrane raft /  lipid binding / lipid binding /  enzyme binding / enzyme binding /  cell surface / cell surface /  ATP hydrolysis activity / protein-containing complex / ATP hydrolysis activity / protein-containing complex /  mitochondrion / mitochondrion /  RNA binding / extracellular exosome / RNA binding / extracellular exosome /  ATP binding / ATP binding /  membrane / identical protein binding / membrane / identical protein binding /  nucleus / nucleus /  plasma membrane plasma membraneSimilarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.89 Å cryo EM / Resolution: 2.89 Å | ||||||||||||
 Authors Authors | Lai, Y. / Zhang, Y. / Gong, H. | ||||||||||||
| Funding support |  China, 3items China, 3items
| ||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of Mycobacterium tuberculosis ATP synthase Authors: Zhang, Y. / Lai, Y. / Liu, F. / Rao, Z. / Gong, H. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8ki3.cif.gz 8ki3.cif.gz | 845.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8ki3.ent.gz pdb8ki3.ent.gz | 696.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8ki3.json.gz 8ki3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ki/8ki3 https://data.pdbj.org/pub/pdb/validation_reports/ki/8ki3 ftp://data.pdbj.org/pub/pdb/validation_reports/ki/8ki3 ftp://data.pdbj.org/pub/pdb/validation_reports/ki/8ki3 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  37251MC 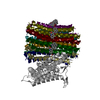 8j57C  8j58C  8jr0C  8jr1C  8khfC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-ATP synthase subunit ... , 12 types, 16 molecules ABCDEFGOHIMNPRST
| #1: Protein |  / ATP synthase F1 subunit alpha / ATP synthase F1 subunit alphaMass: 55276.160 Da / Num. of mol.: 3 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P25705 Homo sapiens (human) / References: UniProt: P25705#2: Protein |  Mass: 51821.965 Da / Num. of mol.: 3 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P06576 Homo sapiens (human) / References: UniProt: P06576#3: Protein | |  / ATP synthase F1 subunit gamma / F-ATPase gamma subunit / ATP synthase F1 subunit gamma / F-ATPase gamma subunitMass: 30207.752 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P36542 Homo sapiens (human) / References: UniProt: P36542#5: Protein | |  / ATP synthase peripheral stalk subunit OSCP / Oligomycin sensitivity conferral protein / OSCP / ATP synthase peripheral stalk subunit OSCP / Oligomycin sensitivity conferral protein / OSCPMass: 20904.488 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P48047 Homo sapiens (human) / References: UniProt: P48047#7: Protein | |  / ATP synthase F1 subunit delta / F-ATPase delta subunit / ATP synthase F1 subunit delta / F-ATPase delta subunitMass: 15029.817 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P30049 Homo sapiens (human) / References: UniProt: P30049#8: Protein | |  / ATPase subunit epsilon / ATP synthase F1 subunit epsilon / ATPase subunit epsilon / ATP synthase F1 subunit epsilonMass: 5790.779 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P56381 Homo sapiens (human) / References: UniProt: P56381#10: Protein | |  / ATPase subunit d / ATP synthase peripheral stalk subunit d / ATPase subunit d / ATP synthase peripheral stalk subunit dMass: 18383.982 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: O75947 Homo sapiens (human) / References: UniProt: O75947#11: Protein | |  / F-ATPase protein 6 / F-ATPase protein 6Mass: 24833.102 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P00846 Homo sapiens (human) / References: UniProt: P00846#12: Protein | |  / 6.8 kDa mitochondrial proteolipid protein / MLQ / ATP synthase membrane subunit 6.8PL / 6.8 kDa mitochondrial proteolipid protein / MLQ / ATP synthase membrane subunit 6.8PLMass: 6673.053 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P56378 Homo sapiens (human) / References: UniProt: P56378#14: Protein | |  / ATP synthase membrane subunit f / ATP synthase membrane subunit fMass: 10804.686 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P56134 Homo sapiens (human) / References: UniProt: P56134#15: Protein | |  / ATPase subunit g / ATP synthase membrane subunit g / ATPase subunit g / ATP synthase membrane subunit gMass: 11309.226 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: O75964 Homo sapiens (human) / References: UniProt: O75964#16: Protein | |  / ATPase subunit e / ATP synthase membrane subunit e / ATPase subunit e / ATP synthase membrane subunit eMass: 7947.215 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P56385 Homo sapiens (human) / References: UniProt: P56385 |
|---|
-Protein , 3 types, 3 molecules JQL
| #4: Protein | Mass: 9540.627 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: Q9UII2 Homo sapiens (human) / References: UniProt: Q9UII2 |
|---|---|
| #13: Protein |  / A6L / F-ATPase subunit 8 / A6L / F-ATPase subunit 8Mass: 8000.634 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P03928 Homo sapiens (human) / References: UniProt: P03928 |
| #17: Protein | Mass: 12606.499 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P18859 Homo sapiens (human) / References: UniProt: P18859 |
-ATP synthase F(0) complex subunit ... , 2 types, 9 molecules 12345678K
| #6: Protein | Mass: 7610.954 Da / Num. of mol.: 8 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P05496 Homo sapiens (human) / References: UniProt: P05496#9: Protein | | Mass: 24658.586 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Homo sapiens (human) / References: UniProt: P24539 Homo sapiens (human) / References: UniProt: P24539 |
|---|
-Non-polymers , 4 types, 11 molecules 






| #18: Chemical |  Adenosine triphosphate Adenosine triphosphate#19: Chemical | ChemComp-MG / #20: Chemical |  Adenosine diphosphate Adenosine diphosphate#21: Chemical | ChemComp-BQ1 / |  Bedaquiline Bedaquiline |
|---|
-Details
| Has ligand of interest | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: human ATP synthase / Type: COMPLEX / Entity ID: #1-#17 / Source: NATURAL / Type: COMPLEX / Entity ID: #1-#17 / Source: NATURAL |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Source (recombinant) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Buffer solution | pH: 7.4 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2400 nm / Nominal defocus min: 1200 nm Bright-field microscopy / Nominal defocus max: 2400 nm / Nominal defocus min: 1200 nm |
| Image recording | Electron dose: 50 e/Å2 / Film or detector model: FEI FALCON IV (4k x 4k) |
- Processing
Processing
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
3D reconstruction | Resolution: 2.89 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 84037 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller















 PDBj
PDBj






