+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8885 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | RNA Pol II complex arrested by a CPD lesion | |||||||||
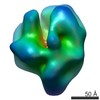
 Map data Map data | RNA Pol II elongation complex with a CPD lesion containing scaffold | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast) Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 10.0 Å cryo EM / Resolution: 10.0 Å | |||||||||
 Authors Authors | Lahiri I / Leschziner AE | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2017 Journal: Nature / Year: 2017Title: Structural basis for the initiation of eukaryotic transcription-coupled DNA repair. Authors: Jun Xu / Indrajit Lahiri / Wei Wang / Adam Wier / Michael A Cianfrocco / Jenny Chong / Alissa A Hare / Peter B Dervan / Frank DiMaio / Andres E Leschziner / Dong Wang /  Abstract: Eukaryotic transcription-coupled repair (TCR) is an important and well-conserved sub-pathway of nucleotide excision repair that preferentially removes DNA lesions from the template strand that block ...Eukaryotic transcription-coupled repair (TCR) is an important and well-conserved sub-pathway of nucleotide excision repair that preferentially removes DNA lesions from the template strand that block translocation of RNA polymerase II (Pol II). Cockayne syndrome group B (CSB, also known as ERCC6) protein in humans (or its yeast orthologues, Rad26 in Saccharomyces cerevisiae and Rhp26 in Schizosaccharomyces pombe) is among the first proteins to be recruited to the lesion-arrested Pol II during the initiation of eukaryotic TCR. Mutations in CSB are associated with the autosomal-recessive neurological disorder Cockayne syndrome, which is characterized by progeriod features, growth failure and photosensitivity. The molecular mechanism of eukaryotic TCR initiation remains unclear, with several long-standing unanswered questions. How cells distinguish DNA lesion-arrested Pol II from other forms of arrested Pol II, the role of CSB in TCR initiation, and how CSB interacts with the arrested Pol II complex are all unknown. The lack of structures of CSB or the Pol II-CSB complex has hindered our ability to address these questions. Here we report the structure of the S. cerevisiae Pol II-Rad26 complex solved by cryo-electron microscopy. The structure reveals that Rad26 binds to the DNA upstream of Pol II, where it markedly alters its path. Our structural and functional data suggest that the conserved Swi2/Snf2-family core ATPase domain promotes the forward movement of Pol II, and elucidate key roles for Rad26 in both TCR and transcription elongation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8885.map.gz emd_8885.map.gz | 4.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8885-v30.xml emd-8885-v30.xml emd-8885.xml emd-8885.xml | 9 KB 9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8885.png emd_8885.png | 37.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8885 http://ftp.pdbj.org/pub/emdb/structures/EMD-8885 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8885 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8885 | HTTPS FTP |
-Related structure data
| Related structure data |  7038C  8735C  8736C  8737C  5vvrC  5vvsC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8885.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8885.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RNA Pol II elongation complex with a CPD lesion containing scaffold | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Binary complex of RNA Pol II and transcription scaffold
| Entire | Name: Binary complex of RNA Pol II and transcription scaffold |
|---|---|
| Components |
|
-Supramolecule #1: Binary complex of RNA Pol II and transcription scaffold
| Supramolecule | Name: Binary complex of RNA Pol II and transcription scaffold type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast) Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast)Strain: ATCC 204508 / S288c |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 7.7 e/Å2 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 10.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: cryoSPARC Details: The particles contributing to the structure adopted a strong preferred orientation. Number images used: 51119 |
 Movie
Movie Controller
Controller