+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Atomic structure of Salmonella SipA/F-actin complex by cryo-EM | |||||||||
 Map data Map data | SipA/F-actin complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  actin / actin /  Salmonella / Salmonella /  type III secretion system / type III secretion system /  SipA / CELL INVASION SipA / CELL INVASION | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoskeletal motor activator activity /  tropomyosin binding / tropomyosin binding /  myosin heavy chain binding / mesenchyme migration / myosin heavy chain binding / mesenchyme migration /  troponin I binding / actin filament bundle / filamentous actin / actin filament bundle assembly / skeletal muscle thin filament assembly / striated muscle thin filament ...cytoskeletal motor activator activity / troponin I binding / actin filament bundle / filamentous actin / actin filament bundle assembly / skeletal muscle thin filament assembly / striated muscle thin filament ...cytoskeletal motor activator activity /  tropomyosin binding / tropomyosin binding /  myosin heavy chain binding / mesenchyme migration / myosin heavy chain binding / mesenchyme migration /  troponin I binding / actin filament bundle / filamentous actin / actin filament bundle assembly / skeletal muscle thin filament assembly / striated muscle thin filament / skeletal muscle myofibril / actin monomer binding / skeletal muscle fiber development / troponin I binding / actin filament bundle / filamentous actin / actin filament bundle assembly / skeletal muscle thin filament assembly / striated muscle thin filament / skeletal muscle myofibril / actin monomer binding / skeletal muscle fiber development /  stress fiber / stress fiber /  titin binding / actin filament polymerization / titin binding / actin filament polymerization /  filopodium / filopodium /  actin filament / actin filament /  Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / calcium-dependent protein binding / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / calcium-dependent protein binding /  lamellipodium / lamellipodium /  cell body / cell body /  actin binding / actin binding /  hydrolase activity / protein domain specific binding / hydrolase activity / protein domain specific binding /  calcium ion binding / positive regulation of gene expression / magnesium ion binding / extracellular region / calcium ion binding / positive regulation of gene expression / magnesium ion binding / extracellular region /  ATP binding / identical protein binding / ATP binding / identical protein binding /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Oryctolagus cuniculus (rabbit) / Oryctolagus cuniculus (rabbit) /   Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) | |||||||||
| Method | helical reconstruction /  cryo EM / Resolution: 3.2 Å cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Niedzialkowska E / Runyan L / Kudryashova E / Kudryashov DS / Egelman EH | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Atomic structure of Salmonella SipA/F-actin complex by cryo-EM Authors: Niedzialkowska E / Runyan L / Kudryashova E / Kudryashov DS / Egelman EH | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42161.map.gz emd_42161.map.gz | 204 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42161-v30.xml emd-42161-v30.xml emd-42161.xml emd-42161.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_42161.png emd_42161.png | 78.7 KB | ||
| Filedesc metadata |  emd-42161.cif.gz emd-42161.cif.gz | 6.6 KB | ||
| Others |  emd_42161_half_map_1.map.gz emd_42161_half_map_1.map.gz emd_42161_half_map_2.map.gz emd_42161_half_map_2.map.gz | 200.6 MB 200.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42161 http://ftp.pdbj.org/pub/emdb/structures/EMD-42161 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42161 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42161 | HTTPS FTP |
-Related structure data
| Related structure data |  8ueeMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_42161.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42161.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SipA/F-actin complex | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_42161_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_42161_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Salmonella SipA/F-actin complex
| Entire | Name: Salmonella SipA/F-actin complex |
|---|---|
| Components |
|
-Supramolecule #1: Salmonella SipA/F-actin complex
| Supramolecule | Name: Salmonella SipA/F-actin complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) |
-Supramolecule #2: Actin, alpha skeletal muscle
| Supramolecule | Name: Actin, alpha skeletal muscle / type: complex / ID: 2 / Parent: 1 |
|---|
-Supramolecule #3: SipA
| Supramolecule | Name: SipA / type: complex / ID: 3 / Parent: 1 |
|---|
-Macromolecule #1: Actin, alpha skeletal muscle
| Macromolecule | Name: Actin, alpha skeletal muscle / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) |
| Molecular weight | Theoretical: 41.875633 KDa |
| Sequence | String: DEDETTALVC DNGSGLVKAG FAGDDAPRAV FPSIVGRPRH QGVMVGMGQK DSYVGDEAQS KRGILTLKYP IE(HIC)GII TNW DDMEKIWHHT FYNELRVAPE EHPTLLTEAP LNPKANREKM TQIMFETFNV PAMYVAIQAV LSLYASGRTT GIVLDSG DG VTHNVPIYEG ...String: DEDETTALVC DNGSGLVKAG FAGDDAPRAV FPSIVGRPRH QGVMVGMGQK DSYVGDEAQS KRGILTLKYP IE(HIC)GII TNW DDMEKIWHHT FYNELRVAPE EHPTLLTEAP LNPKANREKM TQIMFETFNV PAMYVAIQAV LSLYASGRTT GIVLDSG DG VTHNVPIYEG YALPHAIMRL DLAGRDLTDY LMKILTERGY SFVTTAEREI VRDIKEKLCY VALDFENEMA TAASSSSL E KSYELPDGQV ITIGNERFRC PETLFQPSFI GMESAGIHET TYNSIMKCDI DIRKDLYANN VMSGGTTMYP GIADRMQKE ITALAPSTMK IKIIAPPERK YSVWIGGSIL ASLSTFQQMW ITKQEYDEAG PSIVHRKCF UniProtKB:  Actin, alpha skeletal muscle Actin, alpha skeletal muscle |
-Macromolecule #2: Cell invasion protein SipA
| Macromolecule | Name: Cell invasion protein SipA / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) |
| Molecular weight | Theoretical: 28.413857 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: TGETTSFDEV DGVTSKSIIG KPVQATVHGV DDNKQQSQTA EIVNVKPLAS QLAGVENVKT DTLQSDTTVI TGNKAGTTDN DNSQTDKTG PFSGLKFKQN SFLSTVPSVT NMHSMHFDAR ETFLGVIRKA LEPDTSTPFP VRRAFDGLRA EILPNDTIKS A ALKAQCSD ...String: TGETTSFDEV DGVTSKSIIG KPVQATVHGV DDNKQQSQTA EIVNVKPLAS QLAGVENVKT DTLQSDTTVI TGNKAGTTDN DNSQTDKTG PFSGLKFKQN SFLSTVPSVT NMHSMHFDAR ETFLGVIRKA LEPDTSTPFP VRRAFDGLRA EILPNDTIKS A ALKAQCSD IDKHPELKAK METLKEVITH HPQKEKLAEI ALQFAREAGL TRLKGETDYV LSNVLDGLIG DGSWRAGPAY ES YLNKPGV DRVITTVDGL HMQR UniProtKB: Cell invasion protein SipA |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 7 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 7 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: PHOSPHATE ION
| Macromolecule | Name: PHOSPHATE ION / type: ligand / ID: 5 / Number of copies: 7 / Formula: PO4 |
|---|---|
| Molecular weight | Theoretical: 94.971 Da |
| Chemical component information | 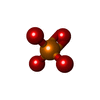 ChemComp-PO4: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 1 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Component - Concentration: 25.0 mM / Component - Formula: TRIS-HCl Tris / Component - Name: TRIS Tris / Component - Name: TRIS Details: Buffer composition: 25 mM TRIS-H-Cl pH 8.0 2 mM DTT |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 291 K / Instrument: LEICA EM GP Details: 3 uL of sample was applied on Lacey grid, then sample was blotted for 3 seconds and plunge-froze in liquid ethane. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.2 µm Bright-field microscopy / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.2 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 12452 / Average exposure time: 5.58 sec. / Average electron dose: 48.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: Details: 8D14 model was used for F-actin 1Q5Z model was used for SipA |
|---|---|
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: cryoSPARC (ver. 4.0.2) / Details: Maximum Likelihood |
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 112.075 Å Applied symmetry - Helical parameters - Δ&Phi: 53.656 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 3.2 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 4.0.2) Details: Reconstruction algorithm: helical refinement and non-uniform refinement in cryoSPARC that is similar to IHRSR algorithm Number images used: 1548993 |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-8uee: |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X


















