+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | AtALMT9 in the apo state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CHANNEL /  TRANSPORT PROTEIN TRANSPORT PROTEIN | |||||||||
| Function / homology | malate transport / Aluminum-activated malate transporter / Aluminium activated malate transporter / plant-type vacuole membrane / monoatomic anion channel activity / Aluminum-activated malate transporter 9 Function and homology information Function and homology information | |||||||||
| Biological species |   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.34 Å cryo EM / Resolution: 3.34 Å | |||||||||
 Authors Authors | Gong DS | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: AtALMT9 in the apo state Authors: Gong DS | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34828.map.gz emd_34828.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34828-v30.xml emd-34828-v30.xml emd-34828.xml emd-34828.xml | 12.7 KB 12.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34828_fsc.xml emd_34828_fsc.xml | 8.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_34828.png emd_34828.png | 231.8 KB | ||
| Filedesc metadata |  emd-34828.cif.gz emd-34828.cif.gz | 5.3 KB | ||
| Others |  emd_34828_half_map_1.map.gz emd_34828_half_map_1.map.gz emd_34828_half_map_2.map.gz emd_34828_half_map_2.map.gz | 59.2 MB 59.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34828 http://ftp.pdbj.org/pub/emdb/structures/EMD-34828 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34828 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34828 | HTTPS FTP |
-Related structure data
| Related structure data |  8hiwMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34828.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34828.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.0742 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34828_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34828_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Dimer of AtALMT9
| Entire | Name: Dimer of AtALMT9 |
|---|---|
| Components |
|
-Supramolecule #1: Dimer of AtALMT9
| Supramolecule | Name: Dimer of AtALMT9 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) |
-Macromolecule #1: Aluminum-activated malate transporter 9
| Macromolecule | Name: Aluminum-activated malate transporter 9 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) |
| Molecular weight | Theoretical: 67.121008 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MAAKQGSFRH GILEKRERLL SNNGFSDFRF TDIESNDLLE NENCGRRTRL CCCCSCGNLS EKISGVYDDA KDVARKAWEM GVSDPRKIV FSAKIGLALT IVALLIFYQE PNPDLSRYSV WAILTVVVVF EFTIGATLSK GFNRALGTLS AGGLALGMAE L STLFGDWE ...String: MAAKQGSFRH GILEKRERLL SNNGFSDFRF TDIESNDLLE NENCGRRTRL CCCCSCGNLS EKISGVYDDA KDVARKAWEM GVSDPRKIV FSAKIGLALT IVALLIFYQE PNPDLSRYSV WAILTVVVVF EFTIGATLSK GFNRALGTLS AGGLALGMAE L STLFGDWE EIFCTLSIFC IGFLATFMKL YPSMKAYEYG FRVFLLTYCY ILISGFRTGQ FIEVAISRFL LIALGAGVSL GV NMFIYPI WAGEDLHNLV VKNFMNVATS LEGCVNGYLR CLEYERIPSK ILTYQASEDP VYKGYRSAVE STSQEESLMS FAI WEPPHG PYKSFNYPWK NYVKLSGALK HCAFTVMALH GCILSEIQAP EERRQVFRQE LQRVGVEGAK LLRELGEKVK KMEK LGPVD LLFEVHLAAE ELQHKIDKKS YLLVNSECWE IGNRATKESE PQELLSLEDS DPPENHAPPI YAFKSLSEAV LEIPP SWGE KNHREALNHR PTFSKQVSWP ARLVLPPHLE TTNGASPLVE TTKTYESASA LSLATFASLL IEFVARLQNV VDAFKE LSQ KANFKEPEIV TTGTDVEFSG ERVGLGQKIR RCFGM UniProtKB: Aluminum-activated malate transporter 9 |
-Macromolecule #2: CITRIC ACID
| Macromolecule | Name: CITRIC ACID / type: ligand / ID: 2 / Number of copies: 1 / Formula: CIT |
|---|---|
| Molecular weight | Theoretical: 192.124 Da |
| Chemical component information | 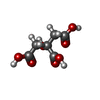 ChemComp-CIT: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.3 µm Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.3 µm |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X


















