+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | ATP- and mtHsp10-bound mtHsp60 V72I | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  chaperonin / chaperonin /  ATPase / ATPase /  foldase / foldase /  CHAPERONE CHAPERONE | |||||||||
| Function / homology |  Function and homology information Function and homology informationcoated vesicle / isotype switching to IgG isotypes /  mitochondrial unfolded protein response / lipopolysaccharide receptor complex / mitochondrial unfolded protein response / lipopolysaccharide receptor complex /  apolipoprotein A-I binding / TFAP2A acts as a transcriptional repressor during retinoic acid induced cell differentiation / protein import into mitochondrial intermembrane space / migrasome / high-density lipoprotein particle binding / positive regulation of T cell mediated immune response to tumor cell ...coated vesicle / isotype switching to IgG isotypes / apolipoprotein A-I binding / TFAP2A acts as a transcriptional repressor during retinoic acid induced cell differentiation / protein import into mitochondrial intermembrane space / migrasome / high-density lipoprotein particle binding / positive regulation of T cell mediated immune response to tumor cell ...coated vesicle / isotype switching to IgG isotypes /  mitochondrial unfolded protein response / lipopolysaccharide receptor complex / mitochondrial unfolded protein response / lipopolysaccharide receptor complex /  apolipoprotein A-I binding / TFAP2A acts as a transcriptional repressor during retinoic acid induced cell differentiation / protein import into mitochondrial intermembrane space / migrasome / high-density lipoprotein particle binding / positive regulation of T cell mediated immune response to tumor cell / apolipoprotein A-I binding / TFAP2A acts as a transcriptional repressor during retinoic acid induced cell differentiation / protein import into mitochondrial intermembrane space / migrasome / high-density lipoprotein particle binding / positive regulation of T cell mediated immune response to tumor cell /  chaperonin ATPase / chaperonin ATPase /  Mitochondrial protein import / positive regulation of macrophage activation / cellular response to interleukin-7 / biological process involved in interaction with symbiont / MyD88-dependent toll-like receptor signaling pathway / 'de novo' protein folding / sperm plasma membrane / B cell activation / Mitochondrial protein import / positive regulation of macrophage activation / cellular response to interleukin-7 / biological process involved in interaction with symbiont / MyD88-dependent toll-like receptor signaling pathway / 'de novo' protein folding / sperm plasma membrane / B cell activation /  DNA replication origin binding / B cell proliferation / apoptotic mitochondrial changes / chaperone cofactor-dependent protein refolding / positive regulation of interferon-alpha production / DNA replication origin binding / B cell proliferation / apoptotic mitochondrial changes / chaperone cofactor-dependent protein refolding / positive regulation of interferon-alpha production /  apolipoprotein binding / positive regulation of interleukin-10 production / protein maturation / RHOG GTPase cycle / response to unfolded protein / chaperone-mediated protein complex assembly / protein folding chaperone / apolipoprotein binding / positive regulation of interleukin-10 production / protein maturation / RHOG GTPase cycle / response to unfolded protein / chaperone-mediated protein complex assembly / protein folding chaperone /  clathrin-coated pit / clathrin-coated pit /  isomerase activity / sperm midpiece / response to cold / positive regulation of interleukin-12 production / isomerase activity / sperm midpiece / response to cold / positive regulation of interleukin-12 production /  T cell activation / T cell activation /  secretory granule / secretory granule /  lipopolysaccharide binding / ATP-dependent protein folding chaperone / osteoblast differentiation / positive regulation of interleukin-6 production / activation of cysteine-type endopeptidase activity involved in apoptotic process / positive regulation of T cell activation / unfolded protein binding / positive regulation of type II interferon production / lipopolysaccharide binding / ATP-dependent protein folding chaperone / osteoblast differentiation / positive regulation of interleukin-6 production / activation of cysteine-type endopeptidase activity involved in apoptotic process / positive regulation of T cell activation / unfolded protein binding / positive regulation of type II interferon production /  p53 binding / p53 binding /  protein folding / protein folding /  single-stranded DNA binding / single-stranded DNA binding /  double-stranded RNA binding / protein-folding chaperone binding / protein refolding / double-stranded RNA binding / protein-folding chaperone binding / protein refolding /  mitochondrial inner membrane / protein stabilization / mitochondrial inner membrane / protein stabilization /  early endosome / early endosome /  mitochondrial matrix / positive regulation of apoptotic process / mitochondrial matrix / positive regulation of apoptotic process /  ubiquitin protein ligase binding / negative regulation of apoptotic process / ubiquitin protein ligase binding / negative regulation of apoptotic process /  enzyme binding / enzyme binding /  cell surface / cell surface /  ATP hydrolysis activity / protein-containing complex / ATP hydrolysis activity / protein-containing complex /  mitochondrion / mitochondrion /  extracellular space / extracellular space /  RNA binding / extracellular exosome / RNA binding / extracellular exosome /  ATP binding / ATP binding /  membrane / membrane /  metal ion binding / metal ion binding /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.7 Å cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Braxton JR / Shao H / Tse E / Gestwicki JE / Southworth DR | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: bioRxiv / Year: 2023 Journal: bioRxiv / Year: 2023Title: Asymmetric apical domain states of mitochondrial Hsp60 coordinate substrate engagement and chaperonin assembly. Authors: Julian R Braxton / Hao Shao / Eric Tse / Jason E Gestwicki / Daniel R Southworth /  Abstract: The mitochondrial chaperonin, mtHsp60, promotes the folding of newly imported and transiently misfolded proteins in the mitochondrial matrix, assisted by its co-chaperone mtHsp10. Despite its ...The mitochondrial chaperonin, mtHsp60, promotes the folding of newly imported and transiently misfolded proteins in the mitochondrial matrix, assisted by its co-chaperone mtHsp10. Despite its essential role in mitochondrial proteostasis, structural insights into how this chaperonin binds to clients and progresses through its ATP-dependent reaction cycle are not clear. Here, we determined cryo-electron microscopy (cryo-EM) structures of a hyperstable disease-associated mtHsp60 mutant, V72I, at three stages in this cycle. Unexpectedly, client density is identified in all states, revealing interactions with mtHsp60's apical domains and C-termini that coordinate client positioning in the folding chamber. We further identify a striking asymmetric arrangement of the apical domains in the ATP state, in which an alternating up/down configuration positions interaction surfaces for simultaneous recruitment of mtHsp10 and client retention. Client is then fully encapsulated in mtHsp60/mtHsp10, revealing prominent contacts at two discrete sites that potentially support maturation. These results identify a new role for the apical domains in coordinating client capture and progression through the cycle, and suggest a conserved mechanism of group I chaperonin function. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29817.map.gz emd_29817.map.gz | 328.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29817-v30.xml emd-29817-v30.xml emd-29817.xml emd-29817.xml | 21.8 KB 21.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29817_fsc.xml emd_29817_fsc.xml emd_29817_fsc_2.xml emd_29817_fsc_2.xml | 14.9 KB 15 KB | Display Display |  FSC data file FSC data file |
| Images |  emd_29817.png emd_29817.png | 177.4 KB | ||
| Others |  emd_29817_additional_1.map.gz emd_29817_additional_1.map.gz emd_29817_half_map_1.map.gz emd_29817_half_map_1.map.gz emd_29817_half_map_2.map.gz emd_29817_half_map_2.map.gz | 173 MB 322.7 MB 322.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29817 http://ftp.pdbj.org/pub/emdb/structures/EMD-29817 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29817 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29817 | HTTPS FTP |
-Related structure data
| Related structure data |  8g7nMC  8g7jC  8g7kC  8g7lC  8g7mC  8g7oC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29817.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29817.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.927 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unsharpened map
| File | emd_29817_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_29817_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_29817_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ATP- and mtHsp10-bound mtHsp60
| Entire | Name: ATP- and mtHsp10-bound mtHsp60 |
|---|---|
| Components |
|
-Supramolecule #1: ATP- and mtHsp10-bound mtHsp60
| Supramolecule | Name: ATP- and mtHsp10-bound mtHsp60 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: 60 kDa heat shock protein, mitochondrial
| Macromolecule | Name: 60 kDa heat shock protein, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 14 / Enantiomer: LEVO / EC number:  chaperonin ATPase chaperonin ATPase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 58.321 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: SNAAKDVKFG ADARALMLQG VDLLADAVAV TMGPKGRTVI IEQSWGSPKV TKDGVTVAKS IDLKDKYKNI GAKLIQDVAN NTNEEAGDG TTTATVLARS IAKEGFEKIS KGANPVEIRR GVMLAVDAVI AELKKQSKPV TTPEEIAQVA TISANGDKEI G NIISDAMK ...String: SNAAKDVKFG ADARALMLQG VDLLADAVAV TMGPKGRTVI IEQSWGSPKV TKDGVTVAKS IDLKDKYKNI GAKLIQDVAN NTNEEAGDG TTTATVLARS IAKEGFEKIS KGANPVEIRR GVMLAVDAVI AELKKQSKPV TTPEEIAQVA TISANGDKEI G NIISDAMK KVGRKGVITV KDGKTLNDEL EIIEGMKFDR GYISPYFINT SKGQKCEFQD AYVLLSEKKI SSIQSIVPAL EI ANAHRKP LVIIAEDVDG EALSTLVLNR LKVGLQVVAV KAPGFGDNRK NQLKDMAIAT GGAVFGEEGL TLNLEDVQPH DLG KVGEVI VTKDDAMLLK GKGDKAQIEK RIQEIIEQLD VTTSEYEKEK LNERLAKLSD GVAVLKVGGT SDVEVNEKKD RVTD ALNAT RAAVEEGIVL GGGCALLRCI PALDSLTPAN EDQKIGIEII KRTLKIPAMT IAKNAGVEGS LIVEKIMQSS SEVGY DAMA GDFVNMVEKG IIDPTKVVRT ALLDAAGVAS LLTTAEVVVT EIPKEEKDPG MGAMGGMGGG MGGGMF UniProtKB: 60 kDa heat shock protein, mitochondrial |
-Macromolecule #2: 10 kDa heat shock protein, mitochondrial
| Macromolecule | Name: 10 kDa heat shock protein, mitochondrial / type: protein_or_peptide / ID: 2 / Number of copies: 14 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.218934 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: SNAMAGQAFR KFLPLFDRVL VERSAAETVT KGGIMLPEKS QGKVLQATVV AVGSGSKGKG GEIQPVSVKV GDKVLLPEYG GTKVVLDDK DYFLFRDGDI LGKYVD UniProtKB: 10 kDa heat shock protein, mitochondrial |
-Macromolecule #3: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 14 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information | 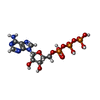 ChemComp-ATP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 14 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 5 / Number of copies: 14 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.8 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 120 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.03 kPa / Details: Pelco easiGlow, 15 mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV / Details: Wait time 10 sec, blot time 3 sec, blot force 0. |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 53937 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 45000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 45000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number grids imaged: 2 / Number real images: 7460 / Average exposure time: 10.0 sec. / Average electron dose: 66.0 e/Å2 |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8g7n: |
 Movie
Movie Controller
Controller












 Z
Z Y
Y X
X


























