+Search query
-Structure paper
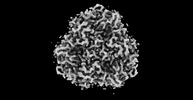
| Title | Structural insights into blue-green light utilization by marine green algal light harvesting complex II at 2.78 Å. |
|---|---|
| Journal, issue, pages | BBA Adv, Vol. 2, Page 100064, Year 2022 |
| Publish date | Nov 11, 2022 |
 Authors Authors | Soichiro Seki / Tetsuko Nakaniwa / Pablo Castro-Hartmann / Kasim Sader / Akihiro Kawamoto / Hideaki Tanaka / Pu Qian / Genji Kurisu / Ritsuko Fujii /   |
| PubMed Abstract | Light-harvesting complex II (LHCII) present in plants and green algae absorbs solar energy to promote photochemical reactions. A marine green macroalga, , exhibits the unique characteristic of ...Light-harvesting complex II (LHCII) present in plants and green algae absorbs solar energy to promote photochemical reactions. A marine green macroalga, , exhibits the unique characteristic of absorbing blue-green light from the sun during photochemical reactions while being underwater owing to the presence of pigment-altered LHCII called siphonaxanthin-chlorophyll binding protein (SCP). In this study, we determined the structure of SCP at a resolution of 2.78 Å using cryogenic electron microscopy. SCP has a trimeric structure, wherein each monomer containing two lutein and two chlorophyll molecules in the plant-type LHCII are replaced by siphonaxanthin and its ester and two chlorophyll molecules, respectively. Siphonaxanthin occupies the binding site in SCP having a polarity in the trimeric inner core, and exhibits a distorted conjugated chain comprising a carbonyl group hydrogen bonded to a cysteine residue of apoprotein. These features suggest that the siphonaxanthin molecule is responsible for the characteristic green absorption of SCP. The replaced chlorophyll molecules extend the region of the stromal side chlorophyll cluster, spanning two adjacent monomers. |
 External links External links |  BBA Adv / BBA Adv /  PubMed:37082593 / PubMed:37082593 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.8 Å |
| Structure data | EMDB-32588, PDB-7wlm: |
| Chemicals |  ChemComp-CHL:  ChemComp-CLA: 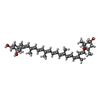 ChemComp-NEX:  ChemComp-0UR:  ChemComp-0IE:  ChemComp-LHG:  ChemComp-HOH: |
| Source |
|
 Keywords Keywords |  PHOTOSYNTHESIS / PHOTOSYNTHESIS /  light-harvesting complex II / siphonaxanthin / Chlorophyll b replacement / blue-green utilization light-harvesting complex II / siphonaxanthin / Chlorophyll b replacement / blue-green utilization |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers