+Search query
-Structure paper
| Title | SARS-CoV-2 can recruit a heme metabolite to evade antibody immunity. |
|---|---|
| Journal, issue, pages | Sci Adv, Vol. 7, Issue 22, Year 2021 |
| Publish date | May 28, 2021 |
 Authors Authors | Annachiara Rosa / Valerie E Pye / Carl Graham / Luke Muir / Jeffrey Seow / Kevin W Ng / Nicola J Cook / Chloe Rees-Spear / Eleanor Parker / Mariana Silva Dos Santos / Carolina Rosadas / Alberto Susana / Hefin Rhys / Andrea Nans / Laura Masino / Chloe Roustan / Evangelos Christodoulou / Rachel Ulferts / Antoni G Wrobel / Charlotte-Eve Short / Michael Fertleman / Rogier W Sanders / Judith Heaney / Moira Spyer / Svend Kjær / Andy Riddell / Michael H Malim / Rupert Beale / James I MacRae / Graham P Taylor / Eleni Nastouli / Marit J van Gils / Peter B Rosenthal / Massimo Pizzato / Myra O McClure / Richard S Tedder / George Kassiotis / Laura E McCoy / Katie J Doores / Peter Cherepanov /     |
| PubMed Abstract | The coronaviral spike is the dominant viral antigen and the target of neutralizing antibodies. We show that SARS-CoV-2 spike binds biliverdin and bilirubin, the tetrapyrrole products of heme ...The coronaviral spike is the dominant viral antigen and the target of neutralizing antibodies. We show that SARS-CoV-2 spike binds biliverdin and bilirubin, the tetrapyrrole products of heme metabolism, with nanomolar affinity. Using cryo-electron microscopy and x-ray crystallography, we mapped the tetrapyrrole interaction pocket to a deep cleft on the spike N-terminal domain (NTD). At physiological concentrations, biliverdin significantly dampened the reactivity of SARS-CoV-2 spike with immune sera and inhibited a subset of neutralizing antibodies. Access to the tetrapyrrole-sensitive epitope is gated by a flexible loop on the distal face of the NTD. Accompanied by profound conformational changes in the NTD, antibody binding requires relocation of the gating loop, which folds into the cleft vacated by the metabolite. Our results indicate that SARS-CoV-2 spike NTD harbors a dominant epitope, access to which can be controlled by an allosteric mechanism that is regulated through recruitment of a metabolite. |
 External links External links |  Sci Adv / Sci Adv /  PubMed:33888467 / PubMed:33888467 /  PubMed Central PubMed Central |
| Methods | EM (single particle) / X-ray diffraction |
| Resolution | 1.82 - 3.6 Å |
| Structure data | EMDB-12585, PDB-7nt9: EMDB-12586, PDB-7nta: EMDB-12587, PDB-7ntc:  PDB-7b62: |
| Chemicals |  ChemComp-BLA:  ChemComp-NAG: 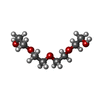 ChemComp-PG4:  ChemComp-PEG: 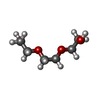 ChemComp-PGE:  ChemComp-1PE:  ChemComp-HOH: |
| Source |
|
 Keywords Keywords |  VIRAL PROTEIN / VIRAL PROTEIN /  SARSCoV2 / SARSCoV2 /  COVID19 / COVID19 /  Biliverdin / Biliverdin /  coronavirus / NTD / green / spike / coronavirus / NTD / green / spike /  glycoprotein / S1 / glycoprotein / S1 /  SARS-CoV-2 / SARS-CoV-2 /  Antibody / Antibody /  Fab / Fab /  epitope epitope |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers











