-検索条件
-検索結果
検索 (著者・登録者: kim & y)の結果1,804件中、1から50件目までを表示しています

EMDB-42676: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

EMDB-42999: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

PDB-8uwl: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

PDB-8v6u: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

EMDB-36271: 
Cryo-EM structure of the DOCK5/ELMO1 complex, focused on one protomer

PDB-8jhk: 
Cryo-EM structure of the DOCK5/ELMO1 complex, focused on one protomer

EMDB-41248: 
Structure of AT118-H Nanobody Antagonist in Complex with the Angiotensin II Type I Receptor

EMDB-41249: 
Structure of AT118-L Nanobody Antagonist in Complex with the Angiotensin II Type I Receptor and Losartan

PDB-8th3: 
Structure of AT118-H Nanobody Antagonist in Complex with the Angiotensin II Type I Receptor

PDB-8th4: 
Structure of AT118-L Nanobody Antagonist in Complex with the Angiotensin II Type I Receptor and Losartan

EMDB-36907: 
Cryo-EM structure of the RC-LH core comples from Halorhodospira halochloris

PDB-8k5o: 
Cryo-EM structure of the RC-LH core comples from Halorhodospira halochloris

EMDB-18658: 
Structure of the NCOA4 (Nuclear Receptor Coactivator 4)-FTH1 (H-Ferritin) complex

PDB-8qu9: 
Structure of the NCOA4 (Nuclear Receptor Coactivator 4)-FTH1 (H-Ferritin) complex

EMDB-36223: 
Cryo-EM structure of the GI.4 Chiba VLP complexed with the CV-1A1 Fv-clasp

PDB-8jg5: 
Cryo-EM structure of the GI.4 Chiba VLP complexed with the CV-1A1 Fv-clasp
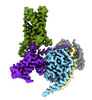
EMDB-43647: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
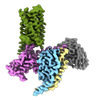
EMDB-43650: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant

EMDB-43656: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)

EMDB-43657: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)

PDB-8vy7: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant

PDB-8vy9: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant

EMDB-43091: 
hGBP1 conformer on the bacterial outer membrane

EMDB-43153: 
hGBP1 conformer on the bacterial outer membrane

EMDB-37386: 
The cryo-EM structure of the Nicotiana tabacum PEP-PAP

EMDB-37387: 
The cryo-EM structure of the Nicotiana tabacum PEP-PAP-TEC1

EMDB-37388: 
The cryo-EM structure of the Nicotiana tabacum PEP-PAP-TEC2

PDB-8w9z: 
The cryo-EM structure of the Nicotiana tabacum PEP-PAP

PDB-8wa0: 
The cryo-EM structure of the Nicotiana tabacum PEP-PAP-TEC1

PDB-8wa1: 
The cryo-EM structure of the Nicotiana tabacum PEP-PAP-TEC2

EMDB-39117: 
Cryo-EM structure of Maltose Binding Protein

PDB-8ybe: 
Cryo-EM structure of Maltose Binding Protein

EMDB-28138: 
3D reconstruction of the apical complex of Plasmodium falciparum (3D7) free merozoite

EMDB-28141: 
3D reconstruction of the apical complex of Plasmodium falciparum (3D7) free merozoite
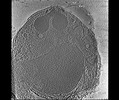
EMDB-28142: 
3D Reconstruction of Plasmodium falciparum (3D7) free merozoite

EMDB-37465: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by sucrose density

EMDB-37466: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by Ca2+-DEAE

PDB-8wdu: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by sucrose density

PDB-8wdv: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by Ca2+-DEAE

EMDB-35377: 
Cryo-EM structure of GPR156 of GPR156-miniGo-scFv16 complex (local refine)

EMDB-35378: 
Cryo-EM structure of miniGo-scFv16 of GPR156-miniGo-scFv16 complex (local refine)

EMDB-35380: 
Cryo-EM structure of GPR156-miniGo-scFv16 complex

EMDB-35382: 
Cryo-EM structure of GPR156A/B of G-protein free GPR156 (local refine)

EMDB-35389: 
Cryo-EM structure of GPR156C/D of G-protein free GPR156 (local refine)

EMDB-35390: 
Cryo-EM structure of G-protein free GPR156

PDB-8ieb: 
Cryo-EM structure of GPR156 of GPR156-miniGo-scFv16 complex (local refine)

PDB-8iec: 
Cryo-EM structure of miniGo-scFv16 of GPR156-miniGo-scFv16 complex (local refine)

PDB-8ied: 
Cryo-EM structure of GPR156-miniGo-scFv16 complex

PDB-8iei: 
Cryo-EM structure of GPR156A/B of G-protein free GPR156 (local refine)

PDB-8iep: 
Cryo-EM structure of GPR156C/D of G-protein free GPR156 (local refine)
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
