-Search query
-Search result
Showing 1 - 50 of 514 items for (author: bian & y)

EMDB-42434: 
Cryo-EM of (L, L)-2NapFF micelle
Method: helical / : Sonani RR, Adams DJ, Egelman EH

EMDB-42436: 
Cryo-EM of (L,D)-2NapFF micelle
Method: helical / : Sonani RR, Adams DJ, Egelman EH

EMDB-18658: 
Structure of the NCOA4 (Nuclear Receptor Coactivator 4)-FTH1 (H-Ferritin) complex
Method: single particle / : Hoelzgen F, Klukin E, Zalk R, Shahar A, Cohen-Schwartz S, Frank GA

PDB-8qu9: 
Structure of the NCOA4 (Nuclear Receptor Coactivator 4)-FTH1 (H-Ferritin) complex
Method: single particle / : Hoelzgen F, Klukin E, Zalk R, Shahar A, Cohen-Schwartz S, Frank GA
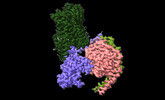
EMDB-42241: 
Serotonin 1E receptor (5-HT1eR)-Gi1 Complex bound with Mianserin
Method: single particle / : Zilberg G, Warren AL, Wacker D

EMDB-42245: 
Serotonin 1E receptor (5-HT1eR)-Gi1 Complex bound with Setiptiline
Method: single particle / : Wacker D, Parpounas AK, Warren AL, Zilberg G

PDB-8ugy: 
Serotonin 1E receptor (5-HT1eR)-Gi1 Complex bound with Mianserin
Method: single particle / : Zilberg G, Warren AL, Wacker D

PDB-8uh3: 
Serotonin 1E receptor (5-HT1eR)-Gi1 Complex bound with Setiptiline
Method: single particle / : Wacker D, Parpounas AK, Warren AL, Zilberg G

EMDB-17924: 
Cryo-EM structure of human Elp123 in complex with tRNA, acetyl-CoA, 5'-deoxyadenosine and methionine
Method: single particle / : Abbassi N, Jaciuk M, Lin TY, Glatt S

EMDB-17925: 
Cryo-EM structure of human Elp123 in complex with 5'-deoxyadenosine and methionine
Method: single particle / : Abbassi N, Jaciuk M, Lin TY, Glatt S

EMDB-17926: 
Cryo-EM structure of human Elp123 in complex with tRNA, S-ethyl-CoA, 5'-deoxyadenosine and methionine
Method: single particle / : Abbassi N, Jaciuk M, Lin TY, Glatt S

EMDB-17927: 
Cryo-EM structure of human Elp123 in complex with tRNA, desulpho-CoA, 5'-deoxyadenosine and methionine
Method: single particle / : Abbassi N, Jaciuk M, Lin TY, Glatt S

PDB-8ptx: 
Cryo-EM structure of human Elp123 in complex with tRNA, acetyl-CoA, 5'-deoxyadenosine and methionine
Method: single particle / : Abbassi N, Jaciuk M, Lin TY, Glatt S

PDB-8pty: 
Cryo-EM structure of human Elp123 in complex with 5'-deoxyadenosine and methionine
Method: single particle / : Abbassi N, Jaciuk M, Lin TY, Glatt S

PDB-8ptz: 
Cryo-EM structure of human Elp123 in complex with tRNA, S-ethyl-CoA, 5'-deoxyadenosine and methionine
Method: single particle / : Abbassi N, Jaciuk M, Lin TY, Glatt S

PDB-8pu0: 
Cryo-EM structure of human Elp123 in complex with tRNA, desulpho-CoA, 5'-deoxyadenosine and methionine
Method: single particle / : Abbassi N, Jaciuk M, Lin TY, Glatt S

EMDB-42023: 
GPR3 Orphan G-coupled Protein Receptor in complex with Dominant Negative Gs.
Method: single particle / : Russell IC, Belousoff MJ, Sexton P

PDB-8u8f: 
GPR3 Orphan G-coupled Protein Receptor in complex with Dominant Negative Gs.
Method: single particle / : Russell IC, Belousoff MJ, Sexton P

EMDB-42371: 
Mouse apoferritin imaged with a square aperture
Method: single particle / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42372: 
Mouse apoferritin imaged with a round aperture
Method: single particle / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42373: 
Mouse apoferritin imaged with a square aperture with P2 projection lens rotation, reconstructed from 25,000 particles
Method: single particle / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42374: 
Mouse apoferritin imaged with a round aperture without P2 projection lens rotation, reconstructed from 25,000 particles
Method: single particle / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42851: 
5x5 tiled montage tomogram of a holey carbon grid with apoferritin imaged with a square electron beam
Method: electron tomography / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42879: 
3x3 tiled montage tomogram of a yeast lamella imaged with a square electron beam
Method: electron tomography / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-18715: 
TDP-43 amyloid fibrils: Morphology-1a
Method: helical / : Sharma K, Shenoy J, Loquet A, Schmidt M, Faendrich M

EMDB-18716: 
TDP-43 amyloid fibrils: Morphology-1b
Method: helical / : Sharma K, Shenoy J, Loquet A, Schmidt M, Faendrich M

EMDB-18717: 
TDP-43 amyloid fibrils: Morphology-2
Method: helical / : Sharma K, Shenoy J, Loquet A, Schmidt M, Faendrich M

PDB-8qx9: 
TDP-43 amyloid fibrils: Morphology-1a
Method: helical / : Sharma K, Shenoy J, Loquet A, Schmidt M, Faendrich M

PDB-8qxa: 
TDP-43 amyloid fibrils: Morphology-1b
Method: helical / : Sharma K, Shenoy J, Loquet A, Schmidt M, Faendrich M

PDB-8qxb: 
TDP-43 amyloid fibrils: Morphology-2
Method: helical / : Sharma K, Shenoy J, Loquet A, Schmidt M, Faendrich M

EMDB-29400: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-29401: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide.
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-29402: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and RG6006
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-29403: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-29404: 
Acinetobacter baylyi LptB2FGC
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-42206: 
Acinetobacter baylyi LptB2FG bound to Acinetobacter baylyi lipopolysaccharide
Method: single particle / : Pahil KS, Gilman MSA, Baidin V, Kruse AC, Kahne D

EMDB-42207: 
Acinetobacter baylyi LptB2FG bound to Acinetobacter baylyi lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Baidin V, Kruse AC, Kahne D

PDB-8frl: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8frm: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide.
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8frn: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and Zosurabalpin
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8fro: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8frp: 
Acinetobacter baylyi LptB2FGC
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8ufg: 
Acinetobacter baylyi LptB2FG bound to Acinetobacter baylyi lipopolysaccharide
Method: single particle / : Pahil KS, Gilman MSA, Baidin V, Kruse AC, Kahne D

PDB-8ufh: 
Acinetobacter baylyi LptB2FG bound to Acinetobacter baylyi lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Baidin V, Kruse AC, Kahne D

EMDB-18941: 
SARS-CoV-2 S (Spike) protein (BA.1) in complex with VHH Ma16B06 (sub-volume of two adjacent RBD-VHH modules)
Method: single particle / : Guttler T, Aksu M, Gorlich D
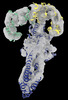
EMDB-18149: 
GBP1 bound by 14-3-3sigma
Method: single particle / : Pfleiderer MM, Liu X, Fisch D, Anastasakou E, Frickel EM, Galej WP

PDB-8q4l: 
GBP1 bound by 14-3-3sigma
Method: single particle / : Pfleiderer MM, Liu X, Fisch D, Anastasakou E, Frickel EM, Galej WP

EMDB-16467: 
Cryo-EM structure of the yeast SPT-Orm1-Monomer complex
Method: single particle / : Schaefer J, Koerner C, Parey K, Januliene D, Moeller A, Froehlich F

EMDB-16468: 
Cryo-EM structure of the yeast SPT-Orm1-Sac1 complex
Method: single particle / : Schaefer J, Koerner C, Parey K, Januliene D, Moeller A, Froehlich F

EMDB-16469: 
Cryo-EM structure of the yeast SPT-Orm1-Dimer complex
Method: single particle / : Schaefer J, Koerner C, Parey K, Januliene D, Moeller A, Froehlich F
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
